
The hardware and bandwidth for this mirror is donated by METANET, the Webhosting and Full Service-Cloud Provider.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]metanet.ch.

fixes is an R package for event study
analysis in panel data — the workhorse tool for visualizing
parallel trends and dynamic treatment effects in
difference-in-differences research.
Version 0.8.0 adds three modern estimators designed for
staggered adoption (where different units adopt
treatment at different times), all accessible through the same
run_es() interface.
Key functions:
| Function | What it does |
|---|---|
run_es() |
Estimate an event study (4 estimators available) |
plot_es() |
Static ggplot2 event study plot |
plot_att_gt() |
Visualize the full ATT(g,t) matrix (CS estimator) |
plot_es_interactive() |
Interactive plotly plot with hover tooltips |
Estimators (selected via the estimator
argument in run_es()):
estimator |
Reference | Best for |
|---|---|---|
"twfe" |
Classic TWFE | Universal treatment timing |
"cs" |
Callaway & Sant’Anna (2021) | Staggered adoption |
"sa" |
Sun & Abraham (2021) | Staggered adoption |
"bjs" |
Borusyak, Jaravel & Spiess (2024) | Staggered adoption |
# From CRAN
install.packages("fixes")
# Development version
pak::pak("yo5uke/fixes")library(fixes)All four estimators share the same interface:
# Classic TWFE (single treatment date)
res_twfe <- run_es(
data = df,
outcome = y,
treatment = treat,
time = period,
timing = 5,
fe = ~ id + period
)
# Sun & Abraham (2021) — staggered adoption
res_sa <- run_es(
data = df_stagg,
outcome = y,
time = year,
timing = timing,
unit = id,
fe = ~ id + year,
staggered = TRUE,
estimator = "sa"
)
# Callaway & Sant'Anna (2021) — staggered adoption
res_cs <- run_es(
data = df_stagg,
outcome = y,
time = year,
timing = timing,
unit = id,
staggered = TRUE,
estimator = "cs",
control_group = "nevertreated"
)
# Borusyak, Jaravel & Spiess (2024) — staggered adoption
res_bjs <- run_es(
data = df_stagg,
outcome = y,
time = year,
timing = timing,
unit = id,
staggered = TRUE,
estimator = "bjs"
)Use run_es() with a fixed event date. Here we use
fixest::base_did, a simulated balanced panel where all
units are treated at period 5.
df <- fixest::base_did
es <- run_es(
data = df,
outcome = y, # outcome variable (unquoted)
treatment = treat, # 0/1 treatment indicator
time = period, # time variable
timing = 5, # treatment occurs at period 5
fe = ~ id + period,
cluster = ~id,
baseline = -1 # period -1 is the reference (default)
)
print(es)## Event Study Result (fixes)
## N: 1080 | Units: NA | Treated units: 1080 | Never-treated: NA
## FE: id + period
## VCOV: HC1 | Cluster: id
## Method: classic | lead_range: 4 lag_range: 5 baseline: -1plot_es(es)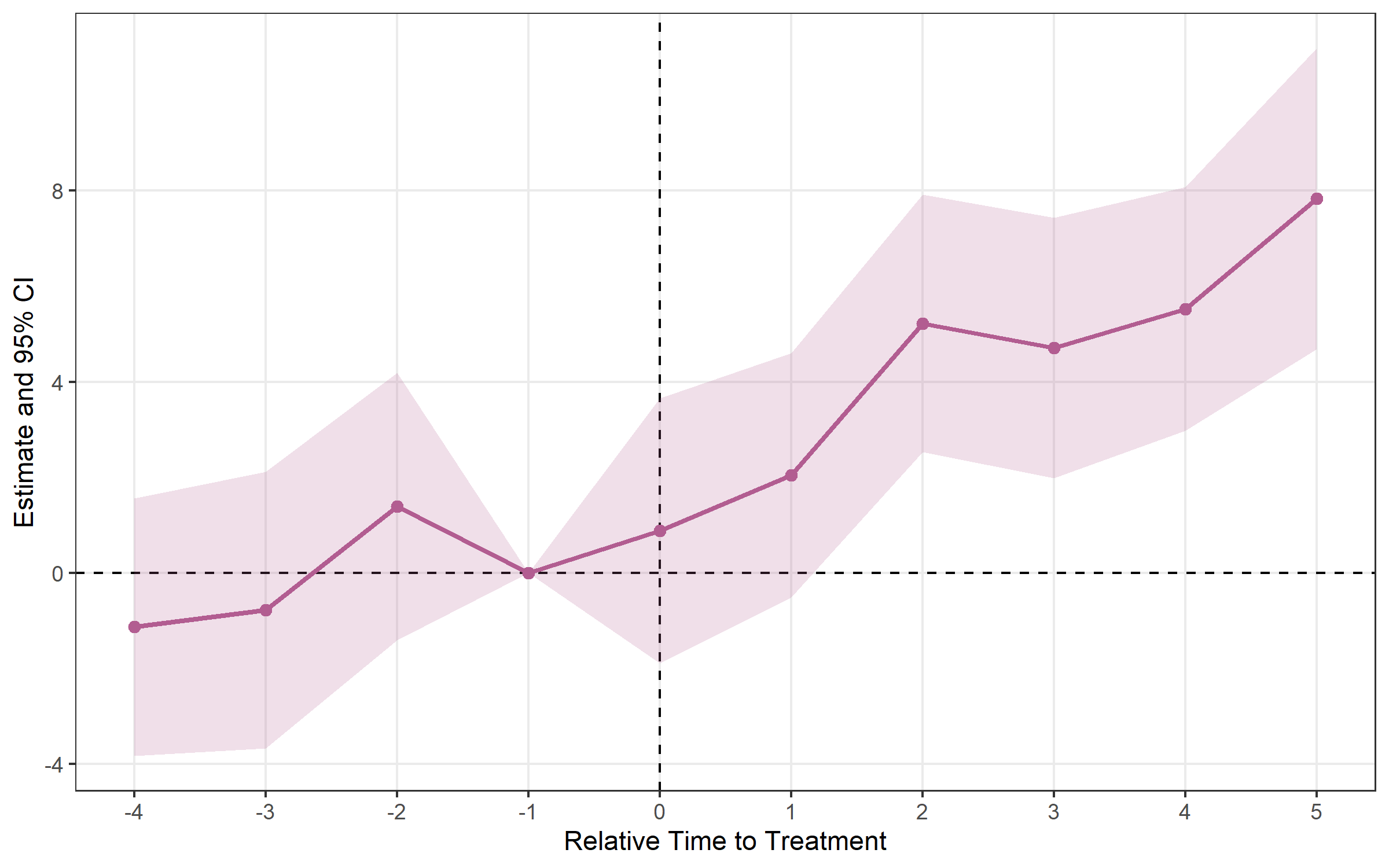
When units adopt treatment at different times, the classic TWFE
estimator can be biased. fixes provides three modern
alternatives.
Setup: create a shared dataset from
fixest::base_stagg. Never-treated units use NA
for their timing column — this is the convention for all three staggered
estimators.
df_stagg <- fixest::base_stagg
df_stagg$timing <- df_stagg$year_treated
df_stagg$timing[df_stagg$year_treated == 10000] <- NA # mark never-treatedestimator = "cs"Estimates a separate ATT(g,t) for every combination of cohort g and calendar time t, then aggregates to the event-study curve. Comparison group can be never-treated units (default) or not-yet-treated units.
cs <- run_es(
data = df_stagg,
outcome = y,
time = year,
timing = timing, # first treatment period; NA = never treated
unit = id,
staggered = TRUE,
estimator = "cs",
control_group = "nevertreated" # or "notyettreated"
)
print(cs)## Event Study Result (fixes)
## N: 950 | Units: 95 | Treated units: 45 | Never-treated: 50
## FE:
## VCOV: analytic | Cluster: -
## Method: classic | lead_range: 9 lag_range: 8 baseline: -1plot_es(cs)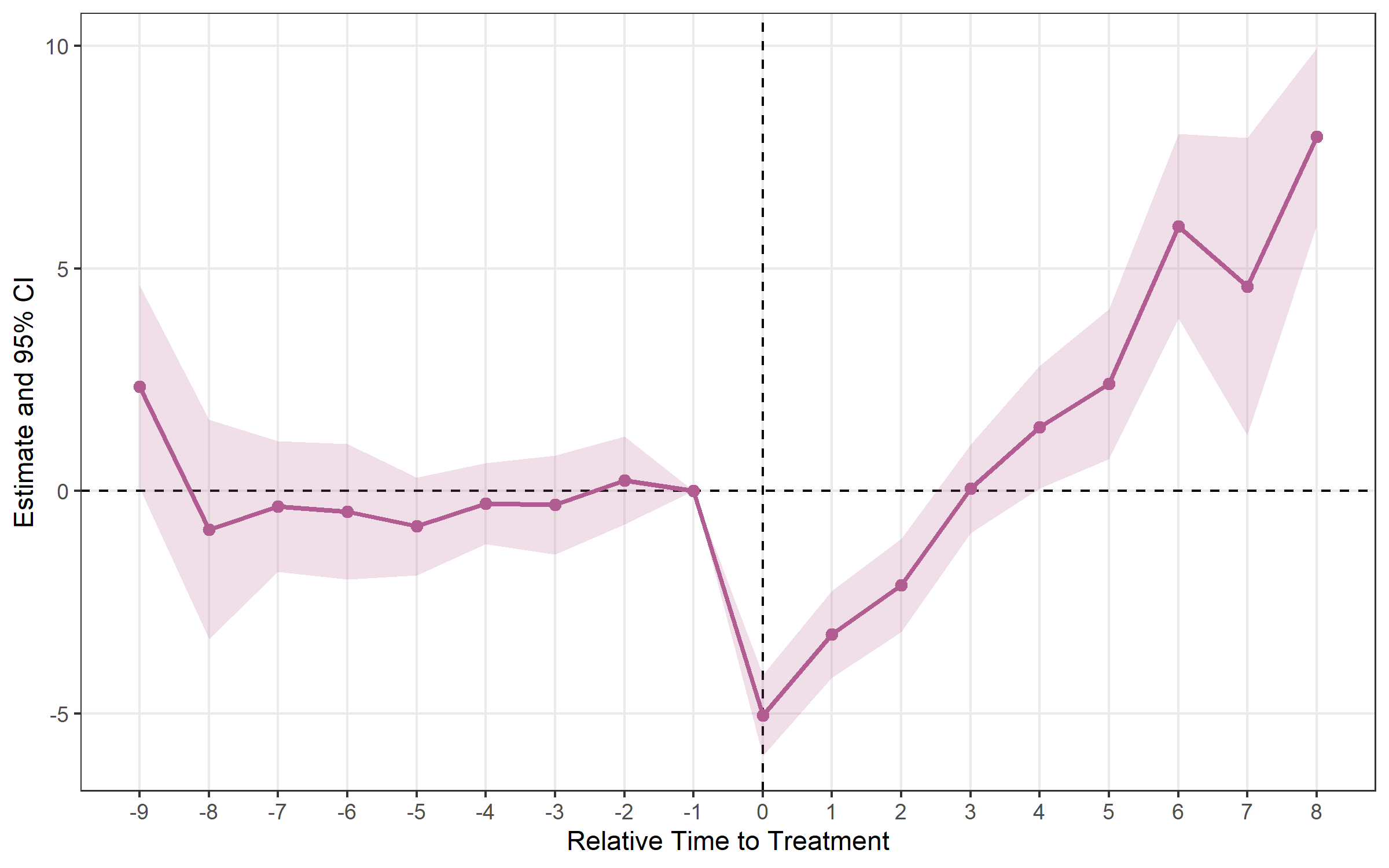
plot_att_gt() shows every cohort × calendar-time cell,
making it easy to spot anticipation effects or heterogeneous
dynamics.
plot_att_gt(cs, type = "heatmap")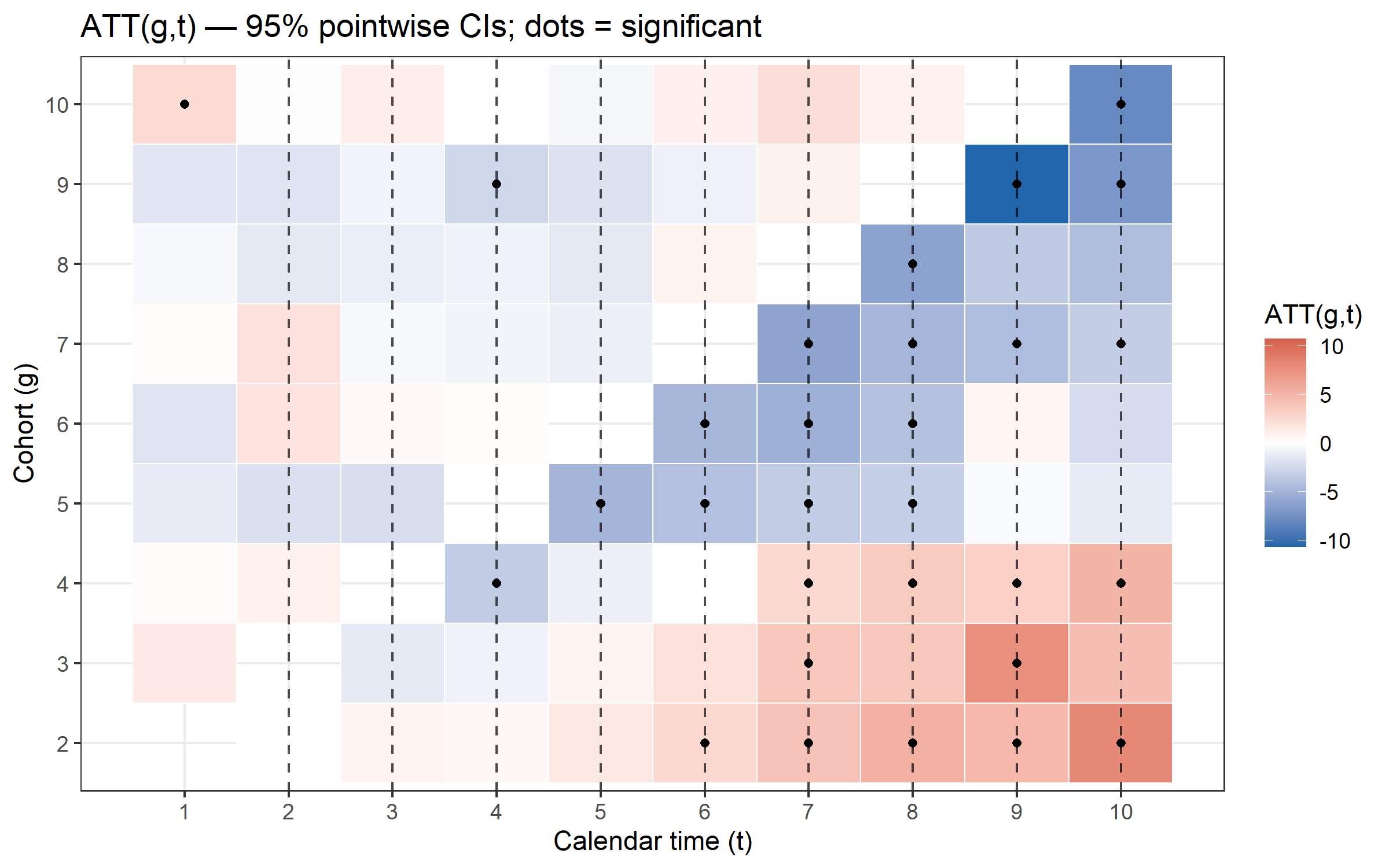
plot_att_gt(cs, type = "facet")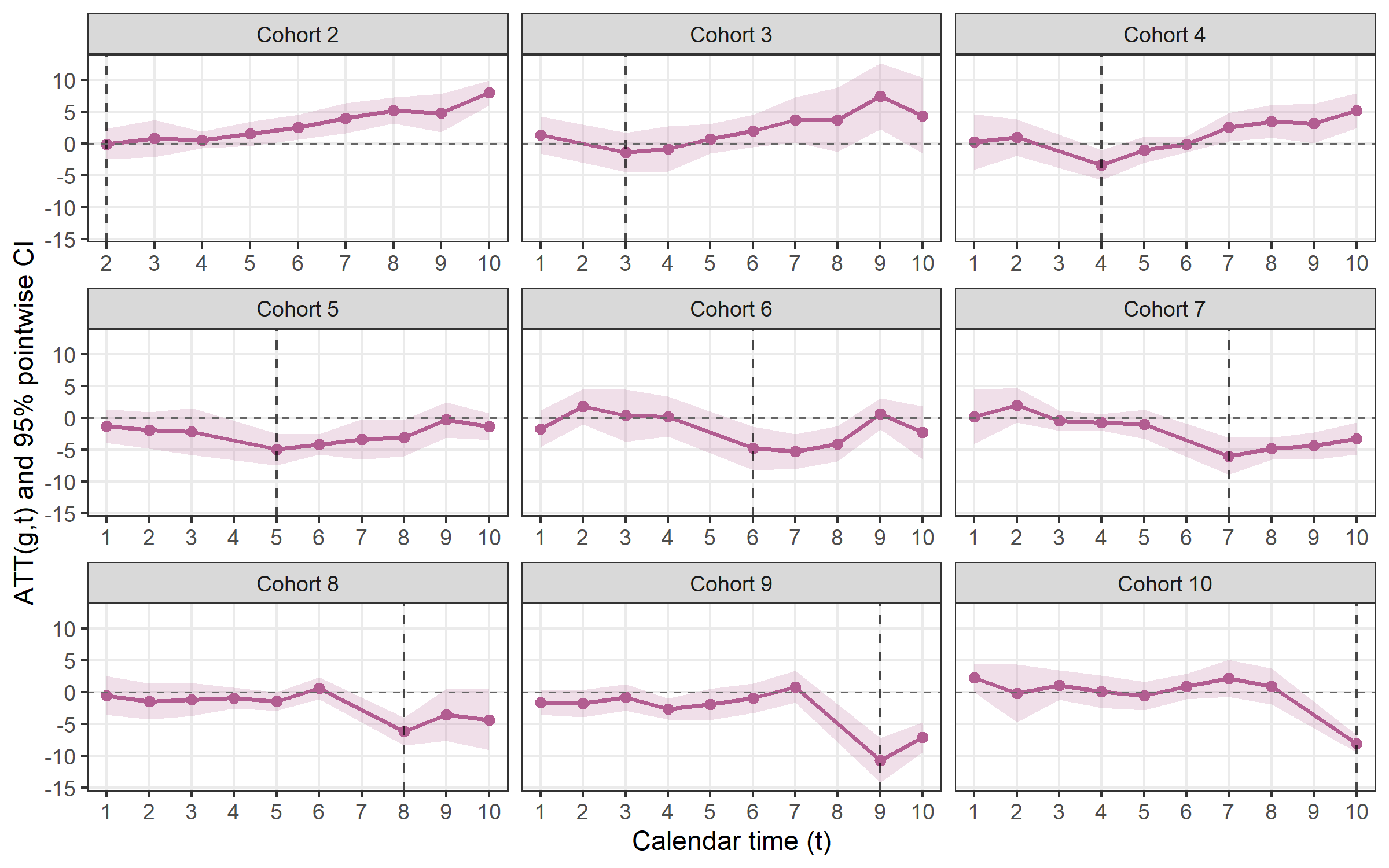
estimator = "sa"Builds cohort × horizon interaction terms, then aggregates with
cohort-share weights. Gives the same result as
fixest::sunab() but through the unified
run_es() interface.
sa <- run_es(
data = df_stagg,
outcome = y,
treatment = treated,
time = year,
timing = timing,
unit = id,
fe = ~ id + year,
staggered = TRUE,
estimator = "sa",
cluster = ~id
)
print(sa)## Event Study Result (fixes)
## N: 950 | Units: 95 | Treated units: 45 | Never-treated: 50
## FE: id + year
## VCOV: HC1 | Cluster: id
## Method: classic | lead_range: 9 lag_range: 8 baseline: -1plot_es(sa)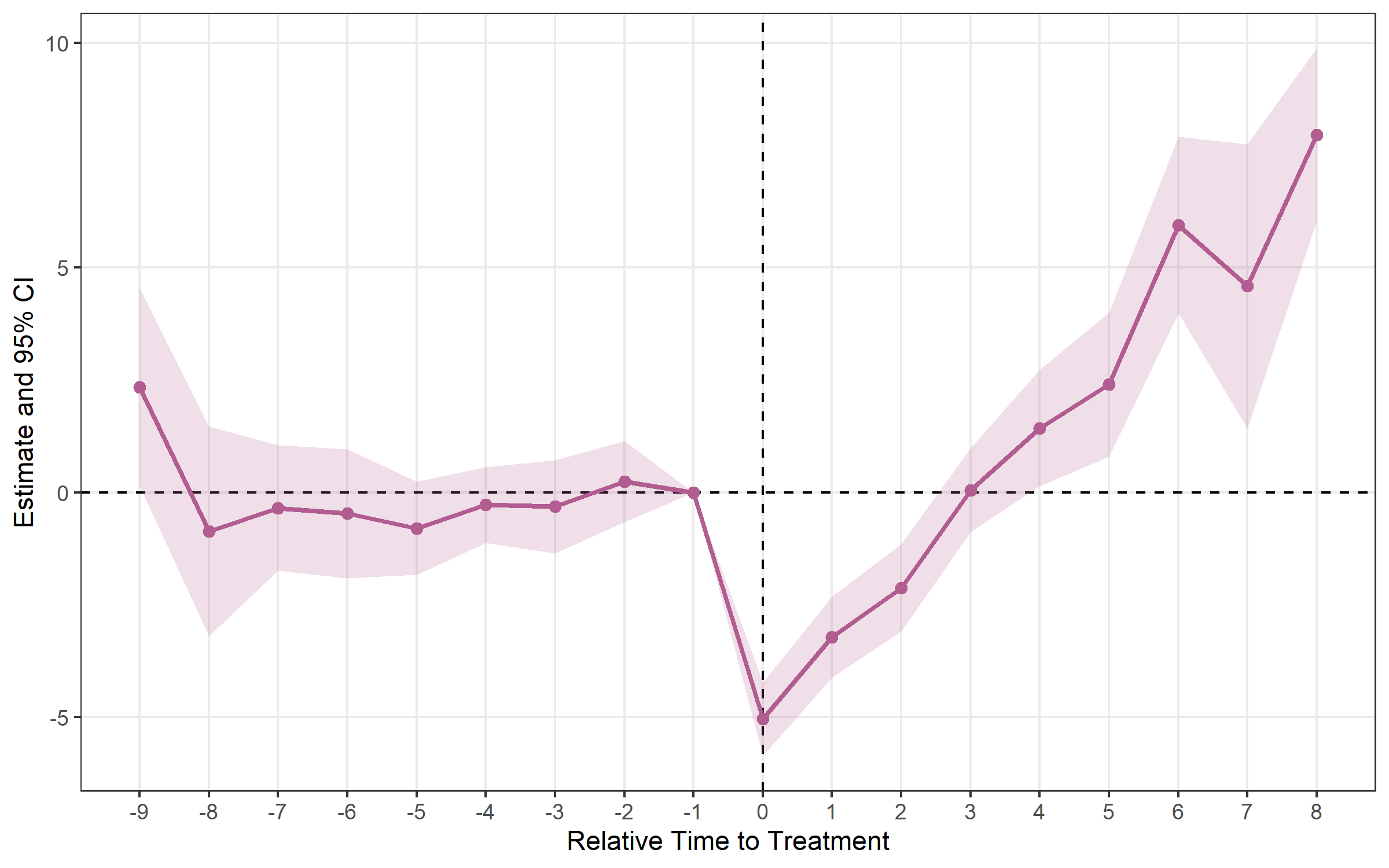
estimator = "bjs"A three-step imputation approach:
bjs <- run_es(
data = df_stagg,
outcome = y,
time = year,
timing = timing,
unit = id,
staggered = TRUE,
estimator = "bjs"
)
print(bjs)## Event Study Result (fixes)
## N: 950 | Units: 95 | Treated units: 45 | Never-treated: 50
## FE: id + year
## VCOV: bjs_conservative | Cluster: -
## Method: classic | lead_range: 1 lag_range: 8 baseline: -1plot_es(bjs)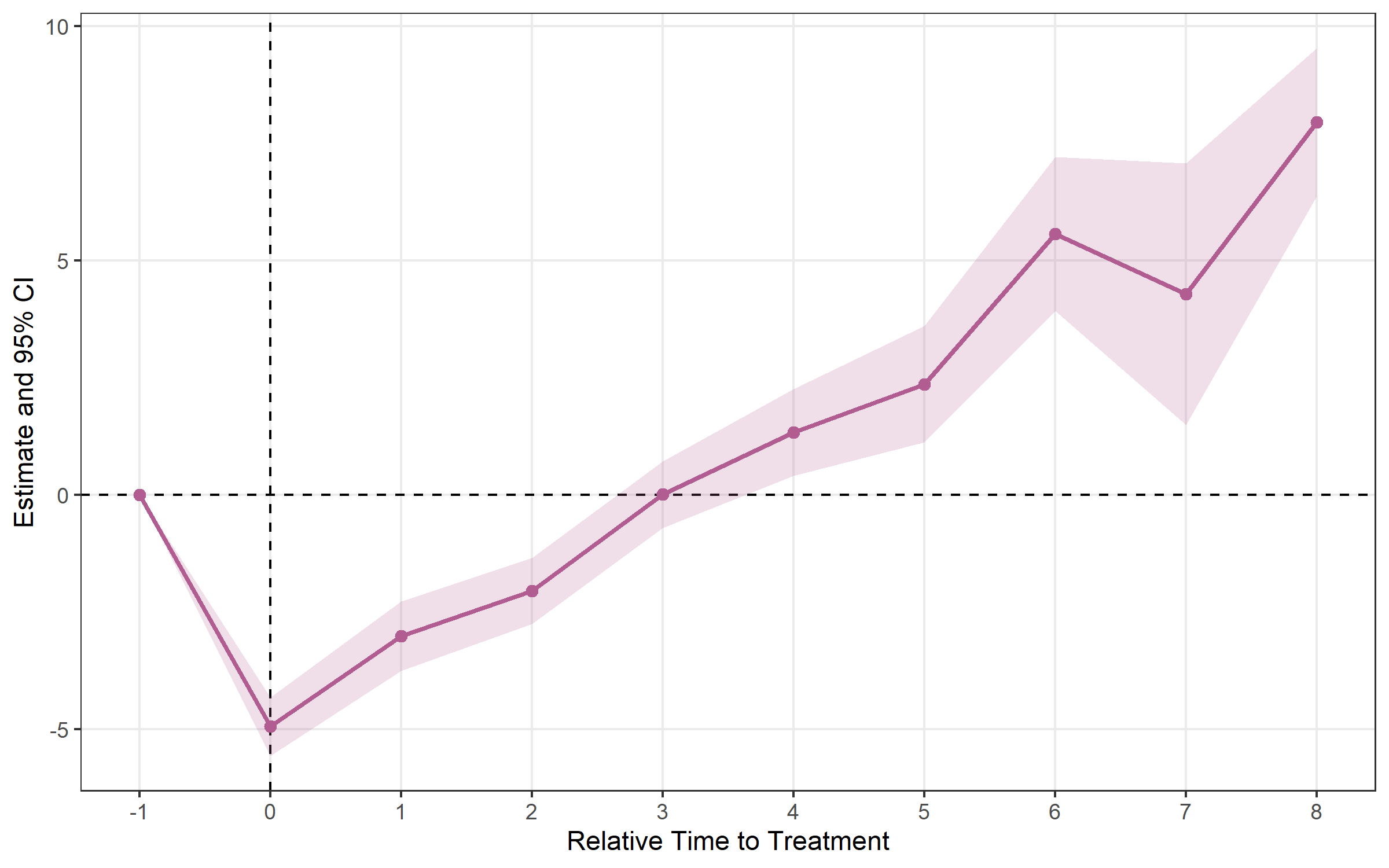
Pointwise confidence intervals control error rates one period at a time. When you display 15 pre- and post-treatment estimates on a single plot, the chance that at least one interval incorrectly excludes zero can be much higher than the nominal 5%.
Simultaneous bands (Callaway & Sant’Anna 2021, Corollary 1) provide joint coverage: with probability ≥ 1 − α, the entire event-study curve is contained within the band.
Use bootstrap = TRUE in run_es() and
show_simultaneous = TRUE in plot_es():
cs_boot <- run_es(
data = df_stagg,
outcome = y,
time = year,
timing = timing,
unit = id,
staggered = TRUE,
estimator = "cs",
control_group = "nevertreated",
bootstrap = TRUE, # run multiplier bootstrap (Algorithm 1, CS 2021)
B = 999, # number of bootstrap draws; 999 recommended in practice
boot_seed = 42 # for reproducibility
)# Lighter outer band = simultaneous CI; darker inner band = pointwise CI
plot_es(cs_boot, show_simultaneous = TRUE)The simultaneous band is always at least as wide as the pointwise band. The interactive plot also gains a second ribbon trace:
plot_es_interactive(cs_boot, show_simultaneous = TRUE)plot_es() works with results from any estimator.
# Error bars instead of ribbon
plot_es(es, type = "errorbar")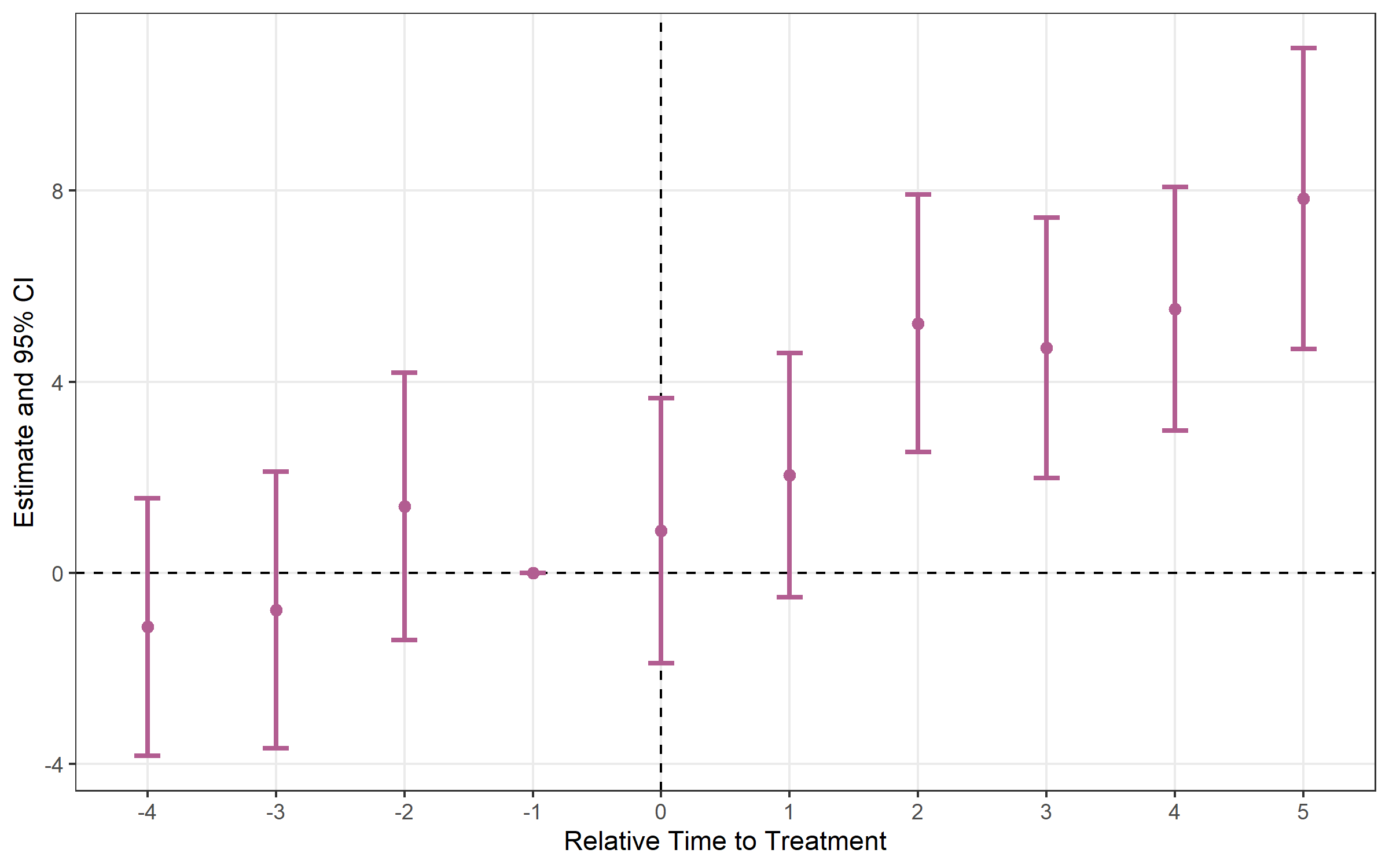
# Multiple CI levels at once
es_multi <- run_es(
data = df,
outcome = y,
treatment = treat,
time = period,
timing = 5,
fe = ~ id + period,
cluster = ~id,
conf.level = c(0.90, 0.95, 0.99)
)
plot_es(es_multi, ci_level = 0.90, theme_style = "minimal")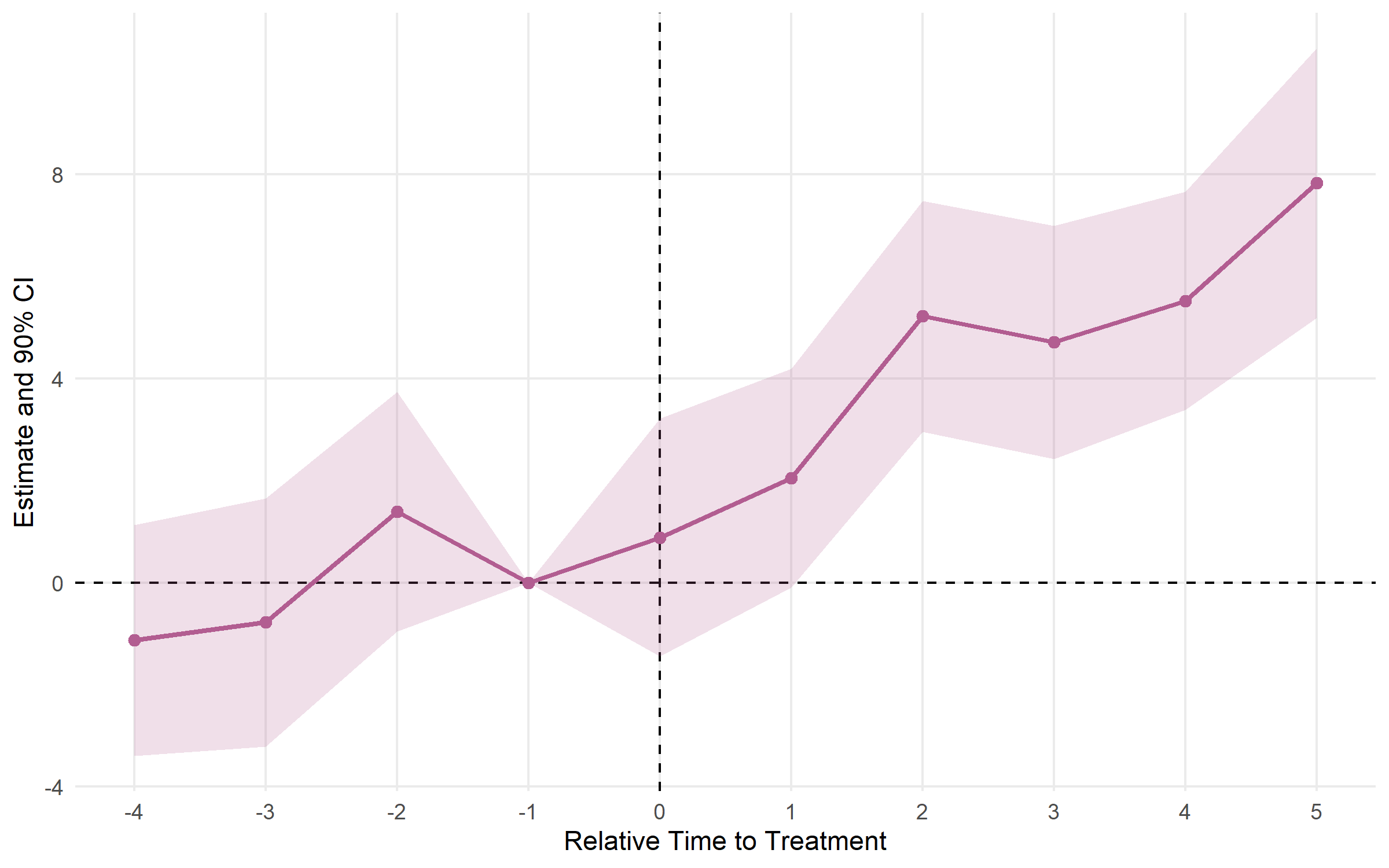
plot_es_interactive() produces a plotly chart with hover
tooltips (requires the plotly package).
plot_es_interactive(es)run_es() arguments| Argument | Default | Description |
|---|---|---|
data |
— | Panel data frame |
outcome |
— | Outcome variable (unquoted) |
treatment |
NULL |
0/1 treatment dummy ("twfe" only) |
time |
— | Time variable (numeric) |
timing |
— | Treatment date (scalar for "twfe", column for others;
NA = never treated) |
unit |
NULL |
Unit ID column (required for "cs", "sa",
"bjs") |
fe |
NULL |
Fixed effects formula, e.g. ~ id + year |
estimator |
"twfe" |
"twfe", "cs", "sa", or
"bjs" |
staggered |
FALSE |
Set TRUE for unit-varying treatment timing |
control_group |
"nevertreated" |
CS only: "nevertreated" or
"notyettreated" |
cluster |
NULL |
Clustering formula, e.g. ~ id |
baseline |
-1 |
Reference period (0 = treatment date) |
lead_range |
auto | Pre-treatment periods to show |
lag_range |
auto | Post-treatment periods to show |
conf.level |
0.95 |
CI level(s); vector allowed, e.g. c(0.90, 0.95) |
vcov |
"HC1" |
VCOV type (any fixest::vcov() type) |
bootstrap |
FALSE |
CS only: run multiplier bootstrap for simultaneous CIs |
B |
999 |
CS bootstrap: number of draws |
boot_seed |
NULL |
CS bootstrap: RNG seed for reproducibility |
Found a bug or have a feature request? Open an issue on GitHub.
These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.