The hardware and bandwidth for this mirror is donated by METANET, the Webhosting and Full Service-Cloud Provider.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]metanet.ch.
Online Time Series Anomaly Detectors
This package provides anomaly detectors in the context of online time series and their evaluation with the Numenta score.
CAD-OSE algorithm is implemented in Python. It uses bencode library in the hashing step. This dependency can be installed with the Python package manager pip.
$ sudo pip install bencode-python3You can install the released version of otsad from CRAN with:
# Get the released version from CRAN
install.packages("otsad")
# Get the latest development version from GitHub
devtools::install_github("alaineiturria/otsad")CpPewmaIpPewmaCpSdEwmaIpSdEwmaCpTsSdEwmaIpTsSdEwmaCpKnnCad(ncm.type = "ICAD")IpKnnCad(ncm.type = "ICAD")CpKnnCad(ncm.type = "LDCD")IpKnnCad(ncm.type = "LDCD")ContextualAnomalyDetectorNormalizeScore +
GetNullAndPerfectScoresReduceAnomaliesPlotDetectionsNOTE: As usual in R, the documentation pages for each function can be loaded from the command line with the commands ? or help:
?CpSdEwma
help(CpSdEwma)This is a basic example of the use of otsad package:
library(otsad)
## basic example code
# Generate data
set.seed(100)
n <- 500
x <- sample(1:100, n, replace = TRUE)
x[70:90] <- sample(110:115, 21, replace = TRUE) # distributional shift
x[25] <- 200 # abrupt transient anomaly
x[320] <- 170 # abrupt transient anomaly
df <- data.frame(timestamp = 1:n, value = x)
# Apply classic processing SD-EWMA detector
result <- CpSdEwma(data = df$value, n.train = 5, threshold = 0.01, l = 3)res <- cbind(df, result)
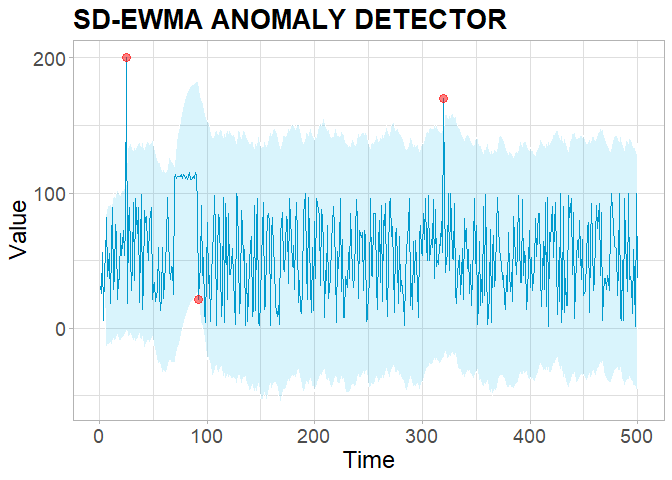
PlotDetections(res, title = "SD-EWMA ANOMALY DETECTOR", return.ggplot = TRUE)
See plotly interactive graph
For more details, see otsad documentation and vignettes.
These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.