The hardware and bandwidth for this mirror is donated by METANET, the Webhosting and Full Service-Cloud Provider.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]metanet.ch.
Nikolas I. Krieger, M.S.;1 Adam T. Perzynski, Ph.D.;2 and Jarrod E. Dalton, Ph.D.1,3
1 Department of Quantitative Health Sciences, Lerner
Research Institute, Cleveland Clinic, 9500 Euclid Avenue (JJN3),
Cleveland, OH, 44195
2 Center for Healthcare Research and Policy, Case Western
Reserve University at MetroHealth, 2500 MetroHealth Drive, Cleveland, OH
44109
3 Cleveland Clinic Lerner College of Medicine, Case Western
Reserve University
Acknowledgements:
The authors of this package acknowledge the support provided by members
of the Northeast Ohio Cohort for Atherosclerotic Risk Estimation
(NEOCARE) investigative team: Claudia Coulton, Douglas Gunzler, Darcy
Freedman, Neal Dawson, Michael Rothberg, David Zidar, David Kaelber,
Douglas Einstadter, Alex Milinovich, Monica Webb Hooper, Kristen
Hassmiller-Lich, Ye Tian (Devin), Kristen Berg, and Sandy Andrukat.
Funding:
This work was supported by The National Institute on Aging of the
National Institutes of Health under award number R01AG055480. The
content is solely the responsibility of the authors and does not
necessarily represent the official views of the National Institutes of
Health.
You can install projects with:
install.packages("projects")The goal of the projects R package is to provide a set
of tools that support an efficient project management workflow for
statisticians and data scientists who perform reproducible research
within team science environments. The projects package is
built upon some existing tools for reproducible research, particularly
RStudio, the R integrated development environment in which it dwells,
and R Markdown, the file structure that allows users to assemble
datasets, to perform analyses and to write manuscripts in a single file.
The projects package is oriented towards efficient and
reproducible academic research manuscript development and incorporates
protocol and analysis templates based on widely-accepted reporting
guidelines (viz., CONSORT and STROBE). When used on a shared file system
(e.g., a server), the projects package provides
infrastructure for collaborative research: multiple researchers can work
on the same project and keep track of its progress without having to
request updates.
The primary features of the projects R package are the following:
At its outset, the projects package creates a folder
called /projects in a user-specified location. This directory
will contain all research projects created and maintained by the user.
The /projects folder will also contain a relational database of
the research projects and the persons who contribute to them. The
database allows for users to input important metadata about the projects
and authors, including stage of research and contact information. Once
this higher-level folder is created, users run R functions to create
projects, each of which is given its own folder. New project folders
automatically include aptly named subfolders and templates in order to
guide the project workflow (e.g., a “data” subfolder; a “datawork” R
Markdown template). Right away, users can begin working on the research
project and edit the metadata of the project itself and its authors. To
lessen the burden of the mundane details of manuscript writing, the
projects package can output lines to the console that, when
copied into an R Markdown file, generates a title page with all relevant
authorship information of any given project. Finally, since users may
create dozens of projects over time, users can run functions to organize
their projects within grouping subfolders of the main /projects
folder.
Reproducibility in research is the focus of the much debated replication crisis and is therefore an increasingly central goal of the contemporary scientific process. In addition to a final report of study results, reproducible research processes include the entire workflow that the researchers used to generate those results. Actively maintaining and archiving this workflow is important to the evaluation and validation of the research. If other researchers can follow the same workflow to achieve the same results, it corroborates the results as scientific knowledge. When results are not produced by the same workflow, however, scientific knowledge is still advanced, as the workflow is shown not to yield generalizable results. (Baker 2016)
There exist today widely available tools that aid with reproducible research, such as R and other statistical programming languages, that allow for precise documentation of some of the most detail-oriented portions of a project workflow. Researchers can distribute their code scripts alongside their results in order to communicate the integrity of their data processing and analysis. Unfortunately, statistical programming languages per se only contribute to research reproducibility insofar as individual statistical programmers are able (1) to use these tools effectively and (2) to integrate their own use of these tools with their collaborators’ work—which may not necessarily be oriented towards reproducibility.
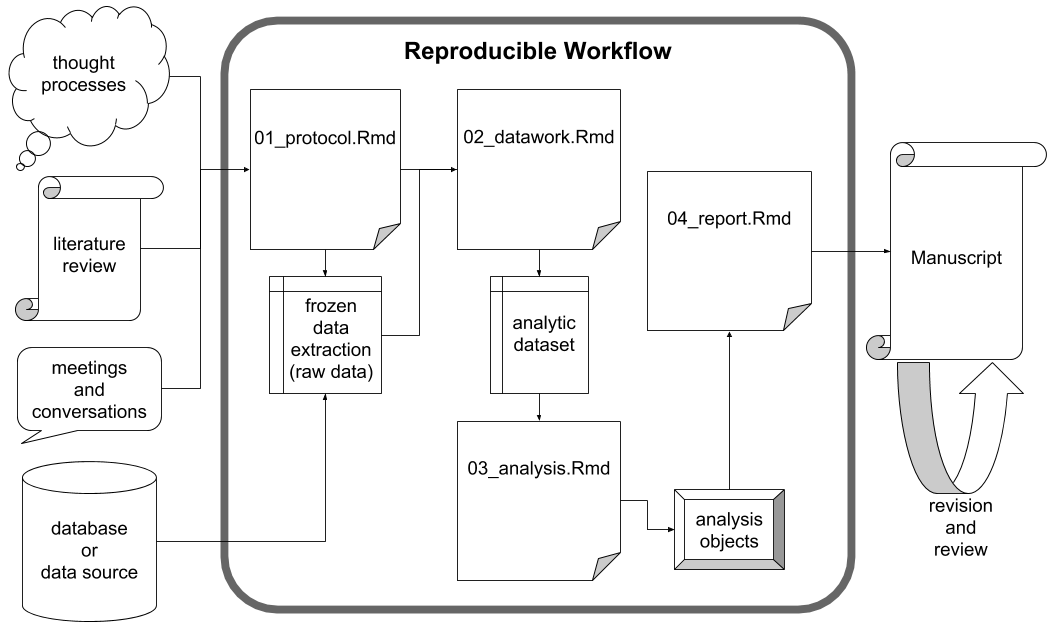
Although researchers of different disciplines may operate in nuanced ways, there are aspects of the project workflow that are common to most investigations. First, studies are conceptualized and designed according to a protocol that details the research questions and planned analyses. Data are collected, manipulated (or “tidied”) in order to make data analysis possible. The results of the analyses are compiled into a report, and ultimately an academic manuscript is drafted and submitted for wider distribution.
When navigating this workflow, researchers strive for reproducibility wherever possible, but especially during the intermediate, data-focused phases of the workflow. Readers of the final manuscript should have access to the study data in its most unrefined state possible: a frozen dataset. A frozen dataset is almost invariably a digital file or set of files that standard data analysis software can process. Whereas study data may have been initially collected in a non-digital manner, a frozen data set represents the study data’s earliest state of simultaneous digitization and consolidation. From this point forward through the reporting stage, total reproducibility is expected. Thanks to modern data analysis via statistical programming languages, a reader should be able to exactly reproduce all data-derived results from the frozen data set alone. With access to the exact scripts the researchers used to produce their results, readers can scrutinize every function call performed on the frozen data set and its descendants.
The middle stage of the assumed study workflow can be performed with near perfect reproducibility, but the beginning and ending stages may not. Researchers cannot document every thought process, literature probe and informal conversation that contributes to the development of the initial study protocol, but they should strive to document it as meticulously as possible. Databases tend to be dynamic such that a given analytic data set is merely a snapshot in time. As for the final stages of project development, journals require that manuscripts adhere to specific and unique stylistic guidelines and that they be digitally submitted with file types that are not independently conducive to reproducibility (e.g., .pdf). For instance, even as RStudio supports the creation of submission-ready documents directly from frozen datasets, the vast majority of project teams include experts who do not use RStudio; therefore, the collaborative manuscript editing process ultimately takes place in an environment (e.g., Microsoft Word) that only supports total reproducibility with extraordinary effort. In light of these realities, researchers must do their best during manuscript creation, keeping the process in reproducible environments for as long as possible and otherwise documenting significant changes and alterations.
projects packageAll projects that the user creates with the projects
package—as well as its infrastructure—reside in a main folder called
/projects. Users need not manually create this directory, and
in fact they are encouraged not to manually manipulate any folders that
the projects package involves. Instead, users run the
function setup_projects(), providing the full file path of
the directory in which the user wants the /projects folder to
reside.
Data about authors, institutional and/or department affiliations and projects are stored in .rds files within the main /project directory, so that the user only needs to enter these details once (unless, for example, a co-author changes their name or affiliations). These data are also used to assemble title pages of reports, with automatically generated author lists and lists of author affiliations. We provide a complete example of this process below in the Demonstration section below.
The main metadata tables accessible to the user are
projects(), authors() and
affiliations(), via functions thusly named. Two additional
tables are internally created to keep track of associations between
authors and projects and between authors and affiliations (see the
Internal Tables section).
id – an identification number, specifically an integer,
unique among the other projects. This number can be used whenever
needing to identify this project within projects package
functions.title – the title of the project. A nonambiguous
substring of title (i.e., a substring that does not match
any other project) can be used whenever needing to identify this project
within projects package functions.short_title – an optional unique nickname for the
project. A nonambiguous substring of short_title (i.e., a
substring that does not match any other project) can be used whenever
needing to identify this project within projects package
functions. This is useful if users cannot remember the long, formal
project title nor the project id.current_owner – the id of the author who
is responsible for taking action in order that work on the project may
proceed further.status – a short description of the status of the
project. For example, it may elaborate on the value of
current_owner and/or stage.deadline_type – a simple description of the meaning of
the date contained in the next field, deadline.deadline – a date indicating some kind of deadline
whose meaning is described in the previous field,
deadline_type.stage – one of seven predefined stages of project
development that the project is currently in:
c("0: idea","1: design","2: data collection","3: analysis","4: manuscript","5: under review","6: accepted")path – the full file path where the project folder is
located.corresp_auth – the id of the author who
should be contacted for any correspondence relating to the project. This
author’s name will be especially marked on automatically generated title
pages for this project, and his or her contact information will be
especially displayed there as well in a “Corresponding Author”
section.creator – the id of the author who
initially created the project, or the value of
Sys.info()["user"] if the author who ran
new_project() did not enter a value.id – an identification number, specifically an integer,
unique among the other authors. This number can be used whenever needing
to identify this author within projects package
functions.given_names – the given name or names of the author. A
nonambiguous substring of given_names (i.e., a substring
that does not match any other author) can be used whenever needing to
identify this author within projects package functions.
This is included in the automatically generated title pages of the
projects associated with this author.last_name – the last name or names of the author. A
nonambiguous substring of last_name (i.e., a substring that
does not match any other author) can be used whenever needing to
identify this author within projects package functions.
This is included after given_names in the automatically
generated title pages of the projects associated with this author.title – the job title of the author.degree – the abbreviation(s) of the author’s academic
degree(s). This is included after last_name in the
automatically generated title pages of the projects associated with this
author.email – the email address of the author. This is
included in the “Corresponding Author” section of the automatically
generated title pages of projects whose corresp_auth field
contains this author.phone – the phone number of the author. This is
included in the “Corresponding Author” section of the automatically
generated title pages of projects whose corresp_auth field
contains this author.id – an identification number, specifically an integer,
unique among the other affiliations. This number can be used whenever
needing to identify this affiliation within projects
package functions.department_name – the department name of the
affiliation. A nonambiguous substring of department_name
(i.e., a substring that does not match any other affiliation) can be
used whenever needing to identify this affiliation within
projects package functions. This is included in the
affiliations section of the automatically generated title page of
projects associated with authors with this affiliation.institution_name – the name of the overall institution
of the affiliation. A nonambiguous substring of
institution_name (i.e., a substring that does not match any
other affiliation) can be used whenever needing to identify this
affiliation within projects package functions. This is
included after department_name in the affiliations section
of the automatically generated title page of projects associated with
authors with this affiliation.address – the address of the affiliation. This is
included after institution_name in the affiliations section
of the automatically generated title page of projects associated with
authors with this affiliation. It is also included in the “Corresponding
Author” section of the title page when a project’s corresponding author
has this affiliation as his or her primary (i.e., first) affiliation
(see the Internal Tables section).In keeping with relational database theory, there are two
.rds files that keep track of the many-to-many relationships
between projects and authors and between authors and affiliations. Each
has two columns, id1 and id2, that contain the
id numbers of these items. Each row of this table describes
an association. Furthermore, the projects package keeps
track of the order in which these associations appear so that the
automatically generated title pages list authors and affiliations in the
correct order. Users are able to run functions to reorder these
associations as needed.
Users create individual project folders with the function
new_project(). By default, the name of each project folder
is of the form pXXXX, where XXXX is the project’s
id padded with 0s on the left side. Its contents are copied
from a template project folder within the .templates directory
in the main /projects folder.
The default project folder template is structured as follows:
The included subfolders serve to organize the project, while the .Rmd files are templates that facilitate the user’s workflow. The style.css and styles.docx files are a cascading style sheets (CSS) file and a Word style template, which allow for custom styling of knitted HTML files and Word documents, respectively. The citations.bib file is an empty BibTeX file. The default 01_protocol.Rmd and 04_report.Rmd files already reference the .bib and style files, easing the implementation of these features for the user.
The goal of the projects package is to provide a
comprehensive set of tools managing project files in a way that is
self-contained in R and independent of the underlying operating system.
On a daily basis, researchers make, move, copy, delete and archive
files. Through the projects package, researchers can
perform all these actions in an organized manner with an automated file
structure. In fact, users are advised not to manipulate the
/projects folder and its content with their operating system,
so that the package does not lose track of these files. Multiuser
application of projects requires a server or an otherwise
shared directory where multiple users can access the /projects
folder. File-managing functions—along with all functions—are
demonstrated below in the Demonstration section.
Upon installation, the projects package must be set up
using setup_projects(). The user is to input the file path
of the directory wherein the /projects folder is to be
located.
library(projects)setup_projects("~")
#> projects folder created at
#> /tmp/RtmpH0SVxn/projects
#>
#> Add affiliations with new_affiliation(),
#> then add authors with new_author(),
#> then create projects with new_project()As the message suggests, it is in the user’s best interest to add affiliations, followed by authors and projects.
new_affiliation(
department_name = "Department of Physics",
institution_name = "University of North Science",
address = "314 Newton Blvd, Springfield CT 06003"
)
#> New affiliation:
#> # A tibble: 1 x 4
#> id department_name institution_name address
#> <int> <chr> <chr> <chr>
#> 1 1 Department of Physi… University of North Sc… 314 Newton Blvd, Springfie…This affiliation has been successfully added to the “affiliations”
table in the projects relational database. Now to create a
few more affiliations:
new_affiliation(
department_name = "Impossibles Investigation Team",
institution_name = "Creekshirebrook Academy of Thinks",
address = "Let Gade 27182, 1566 Copenhagen, Denmark"
)
#> New affiliation:
#> # A tibble: 1 x 4
#> id department_name institution_name address
#> <int> <chr> <chr> <chr>
#> 1 2 Impossibles Investigat… Creekshirebrook Academy… Let Gade 27182, 1566 C…
new_affiliation(
department_name = "Statistical Consulting Unit",
institution_name = "Creekshirebrook Academy of Thinks",
address = "196 Normal Ave, Columbus, OH ",
id = 50
)
#> New affiliation:
#> # A tibble: 1 x 4
#> id department_name institution_name address
#> <int> <chr> <chr> <chr>
#> 1 50 Statistical Consulting… Creekshirebrook Academy o… "196 Normal Ave, Col…Note that we chose a specific id number (50) for the
affiliation called the “Impossibles Investigation Team.”
Now we are ready to add authors to the “authors” table of the
projects database.
new_author(
given_names = "Scott",
last_name = "Bug",
title = "Professor",
affiliations = c(2, "Physics"),
degree = "PhD",
email = "scottbug@imPOSSible.net",
phone = "965-555-5556"
)
#> New author:
#> # A tibble: 1 x 7
#> id last_name given_names title degree email phone
#> <int> <chr> <chr> <chr> <chr> <chr> <chr>
#> 1 1 Bug Scott Professor PhD scottbug@impossible.… 965-555-55…
#>
#> New author's affiliations:
#> # A tibble: 2 x 4
#> affiliation_id department_name institution_name address
#> <int> <chr> <chr> <chr>
#> 1 2 Impossibles Investig… Creekshirebrook Acade… Let Gade 27182, 1…
#> 2 1 Department of Physics University of North S… 314 Newton Blvd, …Notice that in creating associations between Scott Bug and his
affiliations, we were able to enter both the id number of
one of them (2) and a substring of the department_name of
the other (“Physics”). Also notice that the email address was coerced to
be lowercase.
Now we’ll add more authors.
new_author(
given_names = "Marie",
last_name = "Curie",
title = "Chemist",
affiliations = "Unit",
phone = "553-867-5309",
id = 86
)
#> New author:
#> # A tibble: 1 x 7
#> id last_name given_names title degree email phone
#> <int> <chr> <chr> <chr> <chr> <chr> <chr>
#> 1 86 Curie Marie Chemist <NA> <NA> 553-867-5309
#>
#> New author's affiliations:
#> # A tibble: 1 x 4
#> affiliation_id department_name institution_name address
#> <int> <chr> <chr> <chr>
#> 1 50 Statistical Consulti… Creekshirebrook Academ… "196 Normal Ave,…
new_author(
given_names = "George Washington",
last_name = "Carver",
title = "Astrophysicist",
degree = "MA, MPhil, PhD",
affiliations = c(1, 2, 50),
id = 1337
)
#> New author:
#> # A tibble: 1 x 7
#> id last_name given_names title degree email phone
#> <int> <chr> <chr> <chr> <chr> <chr> <chr>
#> 1 1337 Carver George Washington Astrophysicist MA, MPhil, PhD <NA> <NA>
#>
#> New author's affiliations:
#> # A tibble: 3 x 4
#> affiliation_id department_name institution_name address
#> <int> <chr> <chr> <chr>
#> 1 1 Department of Physics University of North … "314 Newton Blvd, …
#> 2 2 Impossibles Investig… Creekshirebrook Acad… "Let Gade 27182, 1…
#> 3 50 Statistical Consulti… Creekshirebrook Acad… "196 Normal Ave, C…
new_author(last_name = "Archimedes", title = "Mathematician")
#> New author:
#> # A tibble: 1 x 7
#> id last_name given_names title degree email phone
#> <int> <chr> <chr> <chr> <chr> <chr> <chr>
#> 1 2 Archimedes <NA> Mathematician <NA> <NA> <NA>
#>
#> New author's affiliations:
#> None.
new_author(
last_name = "Wu",
given_names = "Chien-Shiung",
title = "Physicist",
affiliations = c("of North", "Statistical Consulting"),
degree = "PhD",
email = "wu@WU.wU"
)
#> New author:
#> # A tibble: 1 x 7
#> id last_name given_names title degree email phone
#> <int> <chr> <chr> <chr> <chr> <chr> <chr>
#> 1 3 Wu Chien-Shiung Physicist PhD wu@wu.wu <NA>
#>
#> New author's affiliations:
#> # A tibble: 2 x 4
#> affiliation_id department_name institution_name address
#> <int> <chr> <chr> <chr>
#> 1 1 Department of Physi… University of North S… "314 Newton Blvd, …
#> 2 50 Statistical Consult… Creekshirebrook Acade… "196 Normal Ave, C…Now that some authors and affiliations have been created, we can view these tables:
authors()
#> # A tibble: 5 x 7
#> id last_name given_names title degree email phone
#> <int> <chr> <chr> <chr> <chr> <chr> <chr>
#> 1 1 Bug Scott Professor PhD scottbug@impo… 965-555…
#> 2 2 Archimedes <NA> Mathemati… <NA> <NA> <NA>
#> 3 3 Wu Chien-Shiung Physicist PhD wu@wu.wu <NA>
#> 4 86 Curie Marie Chemist <NA> <NA> 553-867…
#> 5 1337 Carver George Washing… Astrophys… MA, MPhil… <NA> <NA>
affiliations()
#> # A tibble: 3 x 4
#> id department_name institution_name address
#> <int> <chr> <chr> <chr>
#> 1 1 Department of Physics University of North Sci… "314 Newton Blvd, Spri…
#> 2 2 Impossibles Investigat… Creekshirebrook Academy… "Let Gade 27182, 1566 …
#> 3 50 Statistical Consulting… Creekshirebrook Academy… "196 Normal Ave, Colum…Now we will showcase project creation:
new_project(
title = "Achieving Cold Fusion",
short_title = "ACF",
authors = c("Bug", "Chien-Shiung", 86, 1337),
current_owner = "Carver",
corresp_auth = "Bug",
stage = "1: design",
deadline_type = "Pilot study",
deadline = "2020-12-31"
)
#>
#> Project 1 has been created at
#> /tmp/RtmpH0SVxn/projects/p0001
#> # A tibble: 1 x 6
#> id title stage status deadline_type deadline
#> <int> <chr> <prjstg> <chr> <chr> <dttm>
#> 1 1 Achieving Cold F… 1: design just crea… Pilot study 2020-12-31 00:00:00
#>
#> New project's authors:
#> # A tibble: 4 x 7
#> author_id last_name given_names title degree email phone
#> <int> <chr> <chr> <chr> <chr> <chr> <chr>
#> 1 1 Bug Scott Professor PhD scottbug@imp… 965-555…
#> 2 3 Wu Chien-Shiung Physicist PhD wu@wu.wu <NA>
#> 3 86 Curie Marie Chemist <NA> <NA> 553-867…
#> 4 1337 Carver George Washin… Astrophys… MA, MPhi… <NA> <NA>
#> # A tibble: 1 x 3
#> current_owner corresp_auth creator
#> <prjaut> <prjaut> <prjaut>
#> 1 1337: Carver 1: Bug 0: kriegenNotice that since a creator was not specified, this
field was populated with the value of
Sys.info()["user"].
Notice also that the author order given to the authors
argument in new_project() command has been preserved. Also
notice that Scott Bug has been marked as the corresponding author, and
his contact information has been included. When knitted, this will be a
proper title page.
Now a few more projects will be created:
new_project(
title = "Weighing the Crown",
short_title = "Eureka!",
authors = "Archimedes",
current_owner = "Archimedes",
corresp_auth = "Archimedes",
stage = 4
)
#>
#> Project 2 has been created at
#> /tmp/RtmpH0SVxn/projects/p0002
#> # A tibble: 1 x 6
#> id title stage status deadline_type deadline
#> <int> <chr> <prjstg> <chr> <chr> <dttm>
#> 1 2 Weighing the … 4: manuscript just cre… <NA> NA
#>
#> New project's authors:
#> # A tibble: 1 x 7
#> author_id last_name given_names title degree email phone
#> <int> <chr> <chr> <chr> <chr> <chr> <chr>
#> 1 2 Archimedes <NA> Mathematician <NA> <NA> <NA>
#> # A tibble: 1 x 3
#> current_owner corresp_auth creator
#> <prjaut> <prjaut> <prjaut>
#> 1 2: Archimedes 2: Archimedes 0: kriegen
new_project(
title = "How I Learned to Stop Worrying and Love the Bomb",
short_title = "Dr. Strangelove",
authors = c("wu", 1),
creator = "wu",
current_owner = "George",
corresp_auth = "George",
stage = "under review",
deadline_type = "2nd revision",
deadline = "2030-10-8",
id = 1945,
status = "debating leadership changes",
parent_directory = "top_secret",
make_directories = TRUE
)
#>
#> Project 1945 has been created at
#> /tmp/RtmpH0SVxn/projects/top_secret/p1945
#> # A tibble: 1 x 6
#> id title stage status deadline_type deadline
#> <int> <chr> <prjstg> <chr> <chr> <dttm>
#> 1 1945 How I Learn… 5: under review debating… 2nd revision 2030-10-08 00:00:00
#>
#> New project's authors:
#> # A tibble: 3 x 7
#> author_id last_name given_names title degree email phone
#> <int> <chr> <chr> <chr> <chr> <chr> <chr>
#> 1 3 Wu Chien-Shiung Physicist PhD wu@wu.wu <NA>
#> 2 1 Bug Scott Professor PhD scottbug@imp… 965-555…
#> 3 1337 Carver George Washin… Astrophys… MA, MPhi… <NA> <NA>
#> # A tibble: 1 x 3
#> current_owner corresp_auth creator
#> <prjaut> <prjaut> <prjaut>
#> 1 1337: Carver 1337: Carver 3: Wu
new_project(
title = "Understanding Radon",
short_title = "Rn86",
authors = 86,
creator = 86,
corresp_auth = 86,
stage = "3",
status = "Safety procedures"
)
#>
#> Project 3 has been created at
#> /tmp/RtmpH0SVxn/projects/p0003
#> # A tibble: 1 x 6
#> id title stage status deadline_type deadline
#> <int> <chr> <prjstg> <chr> <chr> <dttm>
#> 1 3 Understandin… 3: analysis Safety proc… <NA> NA
#>
#> New project's authors:
#> # A tibble: 1 x 7
#> author_id last_name given_names title degree email phone
#> <int> <chr> <chr> <chr> <chr> <chr> <chr>
#> 1 86 Curie Marie Chemist <NA> <NA> 553-867-5309
#> # A tibble: 1 x 3
#> current_owner corresp_auth creator
#> <prjaut> <prjaut> <prjaut>
#> 1 86: Curie 86: Curie 86: CurieHere is the list of all projects that have been created:
projects()
#> # A tibble: 4 x 5
#> id title current_owner status stage
#> <int> <chr> <prjaut> <chr> <prjstg>
#> 1 1945 How I Learned to Stop Wor… 1337: Carver debating leade… 5: under review
#> 2 2 Weighing the Crown 2: Archimedes just created 4: manuscript
#> 3 3 Understanding Radon 86: Curie Safety procedu… 3: analysis
#> 4 1 Achieving Cold Fusion 1337: Carver just created 1: designProjects, authors, and affiliations can all be edited with their
respective edit_*() functions. For example, we can add to
and remove affiliations from an author with:
edit_author(author = "Bug", affiliations = ~ + 50 - impossibles)When adding or removing affiliations/authors from an author/project,
a one-sided formula is used: it must begin with a tilde
~, and elements are added with + and removed
with -. Elements can be referred to by their
id numbers or their names, as described above.
A formula is also used in the authors argument in
edit_project():
edit_project(
"Cold",
title = "Cold Fusion Is Actually Impossible",
authors = ~ "archi",
stage = "accepted"
)
#>
#> Header has changed. Reprint it with:
#> header(1)Here, the title and stage of the project
have also been edited.
Note that the default behavior when adding elements is to place them
before the last author (unless there was only one author). This occurs
after elements are removed, as specified by any minus
signs (-) in the formula.
The reorder_authors() function allows for the user to
change author order:
reorder_authors(project = "Cold Fusion", "George", "Bug", 86)
#>
#> Header has changed. Reprint it with:
#> header(1)Author order matters when obtaining title page YAML text to paste into a protocol or manuscript .Rmd file. Best practice is to do so late in the writing process, if, for example, the name of an affiliation changes in the middle of a research project:
edit_affiliation(
affiliation = "Impossibles",
department_name = "Pseudoscience Debunking Unit"
)The text to be pasted in the YAML can be obtained for any given
project using header():
header(project = "Cold")
title: "Cold Fusion Is Actually Impossible"
author:
- George Washington Carver, MA, MPhil, PhD;^1,2,3^ Scott Bug, PhD;^1,3^\* Marie Curie;^3^ Chien-Shiung Wu, PhD;^1,3^ and Archimedes
- ^1^ Department of Physics, University of North Science, 314 Newton Blvd, Springfield CT 06003
- ^2^ Pseudoscience Debunking Unit, Creekshirebrook Academy of Thinks, Let Gade 27182, 1566 Copenhagen, Denmark
- ^3^ Statistical Consulting Unit, Creekshirebrook Academy of Thinks, 196 Normal Ave, Columbus, OH
- \* Corresponding author
- 314 Newton Blvd, Springfield CT 06003
- 965-555-5556
- scottbug@impossible.netIn order to organize projects, users can create subdirectories within
the main /projects folder where individual project folders can
dwell. Among the examples above, this has already occurred with the
project with the nickname (i.e., short_title)
“Dr. Strangelove” because on its creation the arguments
path = top_secret and make_directories = TRUE
were included. The latter argument must be TRUE if the
desired path does not already exist. Observe the path
column among the existing projects (including the path
column in projects() output requires the argument
verbose = TRUE):
projects(verbose = TRUE) %>% select(id, short_title, path)
#> # A tibble: 3 x 3
#> id short_title path
#> <int> <chr> <chr>
#> 1 1945 Dr. Strangelove /tmp/RtmpH0SVxn/projects/top_secret/p1945
#> 2 2 Eureka! /tmp/RtmpH0SVxn/projects/p0002
#> 3 3 Rn86 /tmp/RtmpH0SVxn/projects/p0003Users can also create subdirectories with the function
new_project_group():
new_project_group("Greek_studies/ancient_studies")
#>
#> The following directory was created:
#> /tmp/RtmpH0SVxn/projects/Greek_studies/ancient_studiesIf a project has already been created, it can be moved
not with edit_project() but
move_project(). Users can also copy projects using
copy_project(); everything in the copy will be the same
except its id, folder name (which, again, is based on its
id), path (which, again, is based on its
folder name), and the name of its .Rproj file (which has the
same name as the folder name).
move_project("Crown", path = "Greek_studies/ancient_studies")
#> # A tibble: 1 x 11
#> id title short_title current_owner status deadline_type deadline
#> <int> <chr> <chr> <prjaut> <chr> <chr> <dttm>
#> 1 2 Weig… Eureka! 2: Archimedes just … <NA> NA
#> # … with 4 more variables: stage <prjstg>, path <chr>, corresp_auth <prjaut>,
#> # creator <prjaut>
#>
#> Project 2 moved so that its new path is
#> /tmp/RtmpH0SVxn/projects/Greek_studies/ancient_studies/p0002
copy_project(
project_to_copy = "Radon",
path = "dangerous_studies/radioactive_studies/radon_studies",
make_directories = TRUE
)
#> # A tibble: 1 x 11
#> id title short_title current_owner status deadline_type deadline
#> <int> <chr> <chr> <prjaut> <chr> <chr> <dttm>
#> 1 3 Unde… Rn86 86: Curie Safet… <NA> NA
#> # … with 4 more variables: stage <prjstg>, path <chr>, corresp_auth <prjaut>,
#> # creator <prjaut>
#>
#> Project 4 below is a copy of project 3 and is located at
#> /tmp/RtmpH0SVxn/projects/dangerous_studies/radioactive_studies/radon_studies/p0004
#> # A tibble: 1 x 11
#> id title short_title current_owner status deadline_type deadline
#> <int> <chr> <lgl> <prjaut> <chr> <chr> <dttm>
#> 1 4 Unde… NA 86: Curie Safet… <NA> NA
#> # … with 4 more variables: stage <prjstg>, path <chr>, corresp_auth <prjaut>,
#> # creator <prjaut>
#>
#> The .Rproj file
#> /tmp/RtmpH0SVxn/projects/dangerous_studies/radioactive_studies/radon_studies/p0004/p0003.Rproj
#> was renamed to
#> /tmp/RtmpH0SVxn/projects/dangerous_studies/radioactive_studies/radon_studies/p0004/p0004.Rproj
#>
#> Be sure to change all instances of "p0003" to "p0004" as desired
#> (e.g., .bib files and references to them in YAML headers).projects(c("Crown", "Radon"), verbose = TRUE) %>% select(id, title, path)
#> # A tibble: 3 x 3
#> id title path
#> <int> <chr> <chr>
#> 1 2 Weighing the C… /tmp/RtmpH0SVxn/projects/Greek_studies/ancient_studies/…
#> 2 4 Understanding … /tmp/RtmpH0SVxn/projects/dangerous_studies/radioactive_…
#> 3 3 Understanding … /tmp/RtmpH0SVxn/projects/p0003Projects can also be archived; they are moved into a subdirectory called /archive that is at the same level as the project folder (/pXXXX) before it was run. If this /archive folder does not exist, it will be created.
archive_project("Strangelove")
#> # A tibble: 1 x 11
#> id title short_title current_owner status deadline_type deadline
#> <int> <chr> <chr> <prjaut> <chr> <chr> <dttm>
#> 1 1945 How … Dr. Strang… 1337: Carver debat… 2nd revision 2030-10-08 00:00:00
#> # … with 4 more variables: stage <prjstg>, path <chr>, corresp_auth <prjaut>,
#> # creator <prjaut>
#>
#> The above project was archived and has the file path
#> /tmp/RtmpH0SVxn/projects/top_secret/archive/p1945When a project is archived, it is no longer included in
projects() output unless the user sets
archived = TRUE.
projects(verbose = TRUE) %>% select(id, short_title, path)
#> # A tibble: 3 x 3
#> id short_title path
#> <int> <chr> <chr>
#> 1 2 Eureka! /tmp/RtmpH0SVxn/projects/Greek_studies/ancient_studies/p0002
#> 2 4 <NA> /tmp/RtmpH0SVxn/projects/dangerous_studies/radioactive_stud…
#> 3 3 Rn86 /tmp/RtmpH0SVxn/projects/p0003
projects(verbose = TRUE, archived = TRUE) %>% select(id, short_title, path)
#> # A tibble: 4 x 3
#> id short_title path
#> <int> <chr> <chr>
#> 1 1945 Dr. Strangelo… /tmp/RtmpH0SVxn/projects/top_secret/archive/p1945
#> 2 2 Eureka! /tmp/RtmpH0SVxn/projects/Greek_studies/ancient_studies/p…
#> 3 4 <NA> /tmp/RtmpH0SVxn/projects/dangerous_studies/radioactive_s…
#> 4 3 Rn86 /tmp/RtmpH0SVxn/projects/p0003Lastly, affiliations, authors and projects can be deleted with the
delete_*() functions. Deleting an author is complete: doing
so removes the author from the creator,
current_owner and corresp_auth fields of all
projects. Furthermore, deleting a project also deletes the entire
project folder. Use the delete_*() functions with
caution.
delete_affiliation("north science")
#> # A tibble: 1 x 4
#> id department_name institution_name address
#> <int> <chr> <chr> <chr>
#> 1 1 Department of Physi… University of North Sc… 314 Newton Blvd, Springfie…
#> # A tibble: 1 x 4
#> id department_name institution_name address
#> <int> <chr> <chr> <chr>
#> 1 1 Department of Physi… University of North Sc… 314 Newton Blvd, Springfie…
#> The above affiliation was deleted.
delete_author(2)
#> # A tibble: 1 x 7
#> id last_name given_names title degree email phone
#> <int> <chr> <chr> <chr> <chr> <chr> <chr>
#> 1 2 Archimedes <NA> Mathematician <NA> <NA> <NA>
#> # A tibble: 1 x 7
#> id last_name given_names title degree email phone
#> <int> <chr> <chr> <chr> <chr> <chr> <chr>
#> 1 2 Archimedes <NA> Mathematician <NA> <NA> <NA>
#> The above author was deleted.
delete_project("Crown")
#> # A tibble: 1 x 11
#> id title short_title current_owner status deadline_type deadline
#> <int> <chr> <chr> <prjaut> <chr> <chr> <dttm>
#> 1 2 Weig… Eureka! NA just … <NA> NA
#> # … with 4 more variables: stage <prjstg>, path <chr>, corresp_auth <prjaut>,
#> # creator <prjaut>
#> # A tibble: 1 x 11
#> id title short_title current_owner status deadline_type deadline
#> <int> <chr> <chr> <prjaut> <chr> <chr> <dttm>
#> 1 2 Weig… Eureka! NA just … <NA> NA
#> # … with 4 more variables: stage <prjstg>, path <chr>, corresp_auth <prjaut>,
#> # creator <prjaut>
#>
#> The above project was deleted.The projects package provides a comprehensive set of
tools for reproducible team science workflows. Efficiency in project
management, including manuscript development, is facilitated by an
internal database that keeps record of project details as well as team
members’ affiliations and contact information. For manuscripts, title
pages are automatically generated from this database, and a selection of
manuscript outlines compliant with reporting guidelines are available in
R Markdown format. We believe that the projects package may
be useful for teams that manage multiple collaborative research projects
in various stages of development.
Baker, Monya. 2016. “1,500 Scientists Lift the Lid on Reproducibility.” Nature News 533 (7604): 452.
These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.