
The hardware and bandwidth for this mirror is donated by METANET, the Webhosting and Full Service-Cloud Provider.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]metanet.ch.
sfdep builds on the great shoulders of spdep package for spatial dependence. sfdep creates an sf and tidyverse friendly interface to the package as well as introduces new functionality that is not present in spdep. sfdep utilizes list columns extensively to make this interface possible.
Install the released version from CRAN
install.packages("sfdep")You can install the development version of sfdep like so:
remotes::install_github("josiahparry/sfdep")There are three main categories of functionality relating to geometry neighbors, weights, and local indicators of spatial association (LISAs).
The most fundamental usage is to find contiguous neighbors from a
polygon. This is done with st_contiguity() which, by
default creates queen weights. If rook weights are desired, set
queen = FALSE. Additional arguments can be passed to the
underlying spdep::poly2nb() via ....
st_contiguity() creates an object of class nb
as used by spdep.
library(sf)
library(sfdep)
library(dplyr)
# grab geometry
geo <- st_geometry(guerry)
nb <- st_contiguity(geo)
nb
#> Neighbour list object:
#> Number of regions: 85
#> Number of nonzero links: 420
#> Percentage nonzero weights: 5.813149
#> Average number of links: 4.941176We can identify higher order neighbors with st_nb_lag()
and the cumulative higher order neighbors with
st_nb_lag_cumul().
st_nb_lag(nb, 2)
#> Neighbour list object:
#> Number of regions: 85
#> Number of nonzero links: 756
#> Percentage nonzero weights: 10.46367
#> Average number of links: 8.894118
st_nb_lag_cumul(nb, 2)
#> Neighbour list object:
#> Number of regions: 85
#> Number of nonzero links: 1176
#> Percentage nonzero weights: 16.27682
#> Average number of links: 13.83529Other point geometry neighbor functions are st_knn(),
st_dist_band(), st_nb_dists().
Polygon weights are created with st_weights() (which
calls spdep::nb2listw). By default they are row
standardized weights.
wt <- st_weights(nb)
wt[1:2]
#> [[1]]
#> [1] 0.25 0.25 0.25 0.25
#>
#> [[2]]
#> [1] 0.1666667 0.1666667 0.1666667 0.1666667 0.1666667 0.1666667Other point based weights can be created with
st_nb_dists(), st_kernel_weights() and
st_inverse_weights().
LISAs are created from a combination of neighbors and weights and are intended to be used inside of a dplyr pipeline. The below is a worked example of calculating the spatial lag and the local moran.
g <- guerry %>%
mutate(
nb = st_contiguity(geometry),
wt = st_weights(nb)
)Then calculate the spatial lag with st_lag(). Given that
we’ve only modified an sf object, we can visualize this with
ggplot2.
library(ggplot2)
# create spatial lag
g %>%
mutate(crime_pers_lag = st_lag(crime_pers, nb, wt)) %>%
ggplot(aes(fill = crime_pers_lag)) +
geom_sf(lwd = 0.2, color = "black") +
theme_void()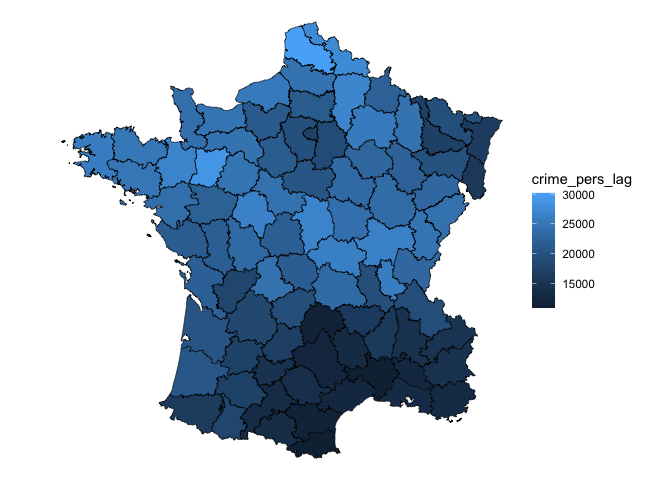
Most users will be interested in local indicators of spatial
association (LISA). Utilize local_moran() to do this.
local_moran() will create a data frame column which
contains a number of informative variables. For example the cluster that
a polygon falls into based on mean, median, or pysal calculations. This
will need to be unnested or certain variables hoisted.
Create the Local Moran data frame column.
lisa <- g %>%
mutate(moran = local_moran(crime_pers, nb, wt))
pull(lisa, moran) %>%
glimpse()
#> Rows: 85
#> Columns: 12
#> $ ii <dbl> 0.52226452, 0.82801651, 0.80353997, 0.74188966, 0.2311871…
#> $ eii <dbl> -0.0436664933, 0.0169987175, -0.0106696690, -0.0015410148…
#> $ var_ii <dbl> 0.3648295427, 0.1244317578, 0.1409560743, 0.2311181704, 0…
#> $ z_ii <dbl> 0.9369545, 2.2991365, 2.1686743, 1.5464057, 1.1463544, 1.…
#> $ p_ii <dbl> 0.348781971, 0.021497187, 0.030107416, 0.122006629, 0.251…
#> $ p_ii_sim <dbl> 0.376, 0.016, 0.036, 0.092, 0.284, 0.124, 0.560, 0.108, 0…
#> $ p_folded_sim <dbl> 0.188, 0.008, 0.018, 0.046, 0.142, 0.062, 0.280, 0.054, 0…
#> $ skewness <dbl> 0.186247324, -0.166050386, -0.065842084, -0.148874532, 0.…
#> $ kurtosis <dbl> -0.256988635, -0.083615702, -0.115769407, -0.105166850, 0…
#> $ mean <fct> High-High, High-High, High-High, Low-Low, Low-Low, Low-Lo…
#> $ median <fct> High-High, High-High, High-High, Low-Low, Low-Low, Low-Lo…
#> $ pysal <fct> High-High, High-High, High-High, Low-Low, Low-Low, Low-Lo…Visualize this by converting insignificant values to NA. This uses a cutoff of 0.1.
lisa %>%
tidyr::unnest(moran) %>%
mutate(pysal = ifelse(p_folded_sim <= 0.1, as.character(pysal), NA)) |>
ggplot(aes(fill = pysal)) +
geom_sf() +
geom_sf(lwd = 0.2, color = "black") +
theme_void() +
scale_fill_manual(values = c("#1C4769", "#24975E", "#EACA97", "#B20016"))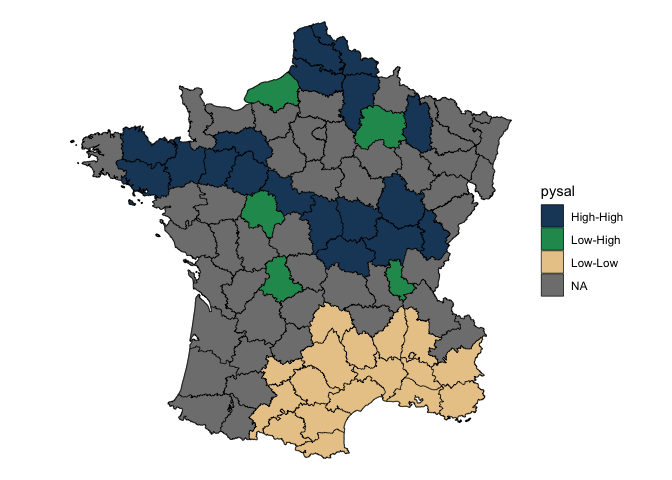
These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.