
The hardware and bandwidth for this mirror is donated by METANET, the Webhosting and Full Service-Cloud Provider.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]metanet.ch.
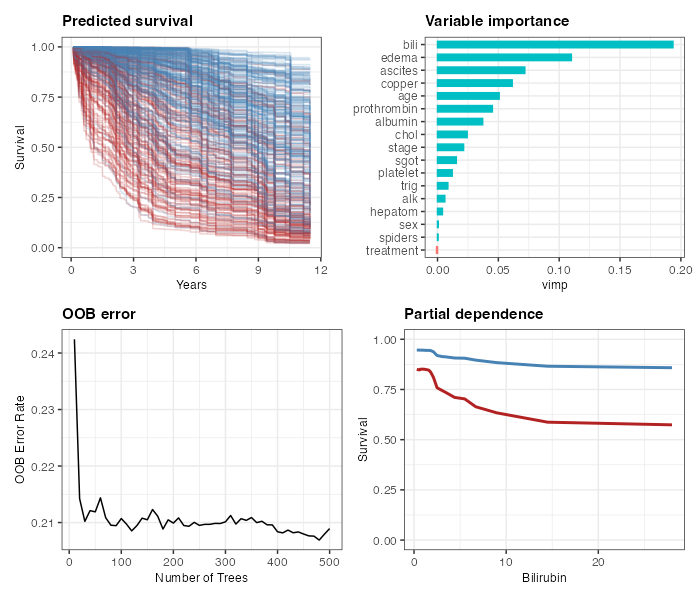
ggRandomForests provides ggplot2-based
diagnostic and exploration plots for random forests fit with randomForestSRC
(>= 3.4.0) or randomForest.
It separates data extraction from plotting so the intermediate tidy
objects can be inspected, saved, or used for custom analyses.
# CRAN (stable)
install.packages("ggRandomForests")
# Development version from GitHub
# install.packages("remotes")
remotes::install_github("ehrlinger/ggRandomForests")library(randomForestSRC)
library(ggRandomForests)
# 1. Fit a forest (regression)
rf <- rfsrc(medv ~ ., data = MASS::Boston, importance = TRUE)
# 2. Check convergence: did the forest grow enough trees?
plot(gg_error(rf))
# 3. Rank predictors by importance
plot(gg_vimp(rf))
# 4. Marginal dependence for top variables
gg_v <- gg_variable(rf)
plot(gg_v, xvar = "lstat")
plot(gg_v, xvar = rf$xvar.names, panel = TRUE, se = FALSE)
# 5. Partial dependence for a single predictor
pv <- plot.variable(rf, xvar.names = "lstat", partial = TRUE, show.plots = FALSE)
pd <- gg_partial(pv)
plot(pd)For survival forests, see the package vignette:
vignette("ggRandomForests")| Function | Input | What you get |
|---|---|---|
gg_error() |
rfsrc / randomForest |
OOB error vs. number of trees |
gg_vimp() |
rfsrc / randomForest |
Variable importance ranking |
gg_rfsrc() |
rfsrc / randomForest |
Predicted vs. observed values |
gg_variable() |
rfsrc / randomForest |
Marginal dependence data frame |
gg_partial() |
plot.variable output |
Partial dependence (continuous + categorical) |
gg_partial_rfsrc() |
rfsrc model |
Partial dependence via partial.rfsrc |
gg_survival() |
rfsrc survival forest |
Kaplan–Meier / Nelson–Aalen estimates |
gg_roc() |
rfsrc / randomForest (class) |
ROC curve data |
gg_brier() |
rfsrc (survival) |
Time-resolved Brier score and CRPS |
Each gg_* function has a corresponding
plot() S3 method that returns a ggplot2
object, making it easy to apply additional ggplot2 layers
or themes. Every gg_* object also implements
print() (header-only summary at the REPL — use
head() for rows) and summary() (printable
diagnostics object).
gg_*
functions extract tidy data objects from the forest. plot()
methods turn those into ggplot2 figures. You can inspect,
save, or transform the data before plotting.ggplot2 composability. Every
plot() method returns a ggplot object that
accepts additional layers, scales, and themes.See NEWS.md for the full changelog. Highlights since v2.4.0:
gg_partial for categorical variables.hvtiRutilities.gg_partial_rfsrc() computes
partial dependence directly from an rfsrc model without a
separate plot.variable call; supports a grouping variable
via xvar2.name.Breiman, L. (2001). Random forests, Machine Learning, 45:5–32.
Ishwaran H. and Kogalur U.B. randomForestSRC: Random Forests for Survival, Regression and Classification. R package version >= 3.4.0. https://cran.r-project.org/package=randomForestSRC
Ishwaran H. and Kogalur U.B. (2007). Random survival forests for R. R News 7(2), 25–31.
Ishwaran H., Kogalur U.B., Blackstone E.H. and Lauer M.S. (2008). Random survival forests. Ann. Appl. Statist. 2(3), 841–860.
Liaw A. and Wiener M. (2002). Classification and Regression by randomForest. R News 2(3), 18–22.
Wickham H. (2009). ggplot2: Elegant Graphics for Data Analysis. Springer New York.
These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.