



The hardware and bandwidth for this mirror is donated by METANET, the Webhosting and Full Service-Cloud Provider.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]metanet.ch.

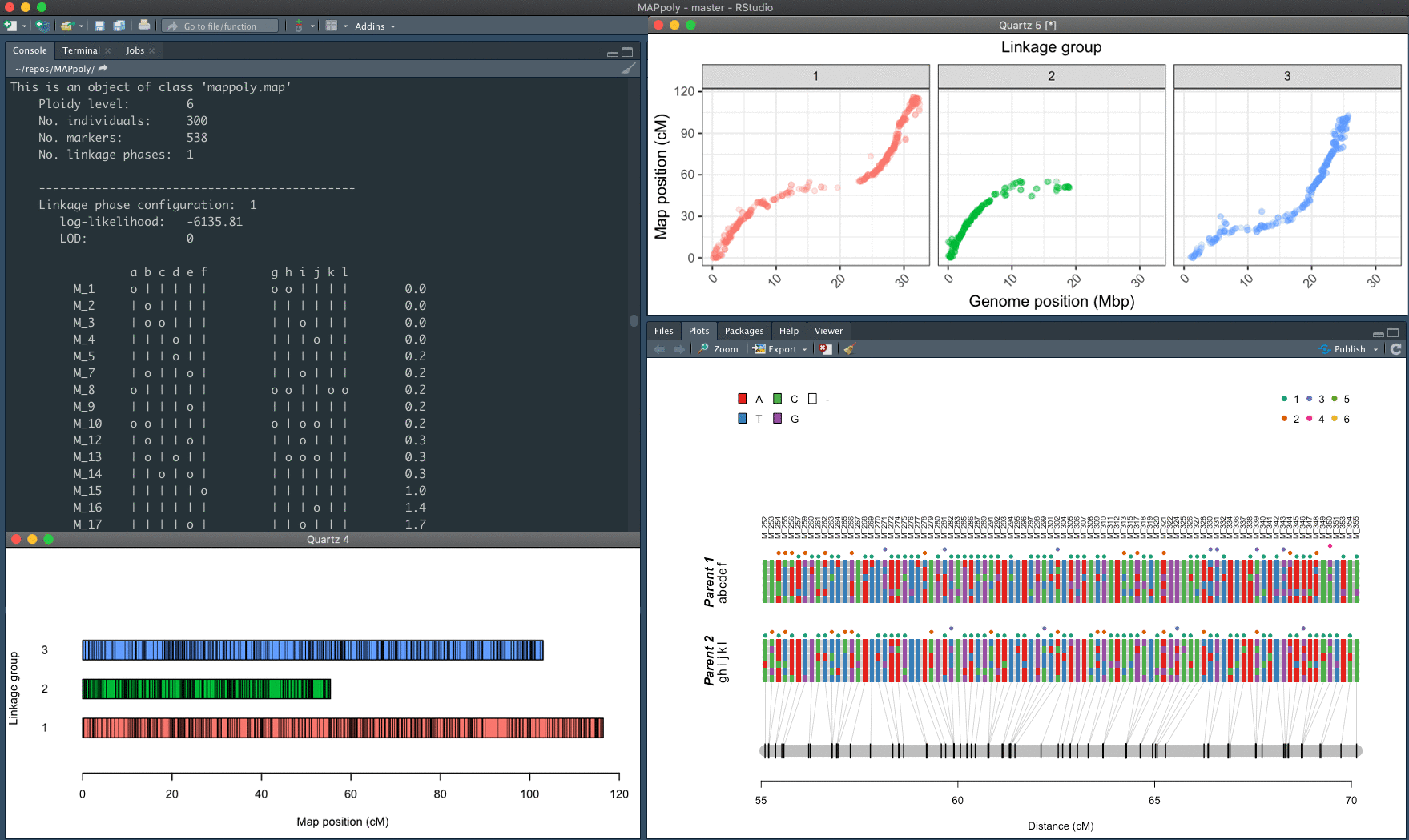
MAPpoly (v. 0.4.1) is an R package to construct genetic maps in diploids and autopolyploids with even ploidy levels. In its current version, MAPpoly can handle ploidy levels up to 8 when using hidden Markov models (HMM) and up to 12 when using the two-point simplification. When dealing with large numbers of markers (> 10,000), we strongly recommend using high-performance computing (HPC).
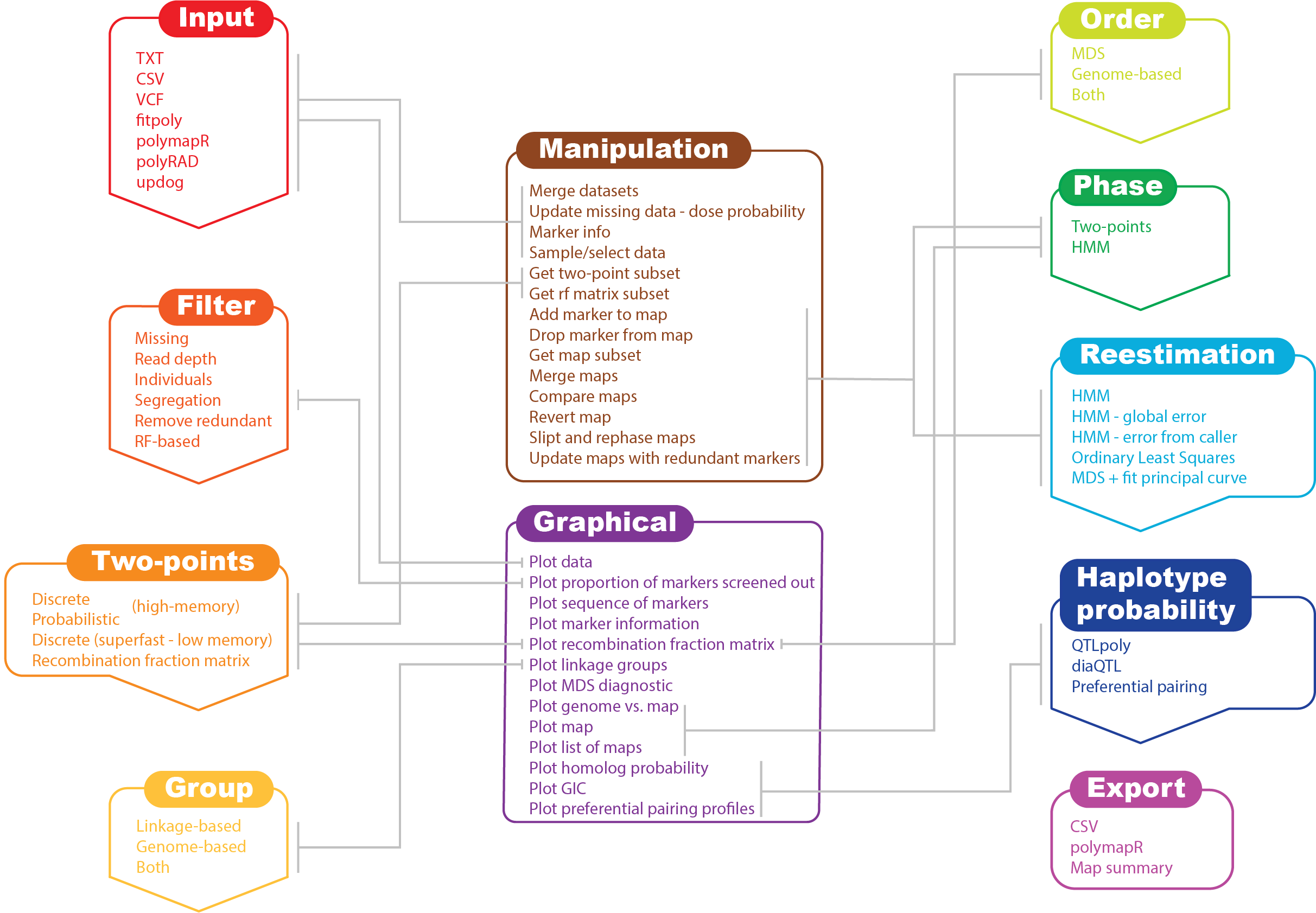
In its current version, MAPpoly can handle the following types of datasets:
MAPpoly also is capable of importing objects generated by the following R packages
The mapping strategy uses pairwise recombination fraction estimation as the first source of information to position allelic variants in specific homologs sequentially. The algorithm relies on the multilocus likelihood obtained through a hidden Markov model (HMM) for situations where pairwise analysis has limited power. The derivation of the HMM used in MAPpoly can be found in Mollinari and Garcia, 2019. The computation of the offspring’s genotypes probabilities and haplotype reconstruction, as well as the preferential pairing profiles, is presented in Mollinari et al., 2020.
To install MAPpoly from the The Comprehensive R Archive Network (CRAN) use
install.packages("mappoly")You can install the development version from Git Hub. Within R, you
need to install pak:
install.packages("pak")If you are using Windows, please install the the latest recommended version of Rtools.
To install MAPpoly from Git Hub use
pak::pak("mmollina/mappoly")For further QTL analysis, we recommend our QTLpoly package. QTLpoly performs random-effect multiple interval mapping (REMIM) in full-sib families of autopolyploid species based on restricted maximum likelihood (REML) estimation and score statistics, as described in Pereira et al. 2020.
We recently released VIEWpoly. VIEWpoly provides a graphical user interface to integrate, visualize and explore results from linkage and quantitative trait loci analysis, together with genomic information for autopolyploid (and diploid) species. The app is meant for interactive use and allows users to optionally upload different sources of information, including gene annotation and alignment files, enabling the exploitation and search for candidate genes in a genome browser. VIEWpoly supports inputs other than MAPpoly’s, including polymapR, diaQTL, QTLpoly, and polyqtlR.
[ ]
]
R vignette("mappoly_startguide")# Enable this universe
options(repos = c(
polyploids = 'https://polyploids.r-universe.dev',
CRAN = 'https://cloud.r-project.org'))
# Install some packages
install.packages('mappoly')This package was developed with support from the Genomic Tools for Sweetpotato Improvement (GT4SP) and SweetGAINS projects, funded by the Bill & Melinda Gates Foundation, and from USDA–NIFA awards including AFRI: “A Genetics-Based Data Analysis System for Breeders in Polyploid Breeding Programs” and SCRI: “Tools for Polyploids.”
These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.