The hardware and bandwidth for this mirror is donated by METANET, the Webhosting and Full Service-Cloud Provider.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]metanet.ch.
The goal of the svyROC package is to plot weighted estimates of the ROC curves and to obtain weighted estimates of the AUC.
The following functions are available:
wse, wsp: estimate sensitivity and
specificity parameters for a specific cut-off point considering sampling
weights.wroc: estimate the ROC curve considering sampling
weights.wauc: estimate the AUC considering sampling
weights.corrected.wauc: correct the optimism of the weighted
estimate of the AUC by means of replicate weights.wocp: calculate optimal cut-off points for individual
classification considering sampling weights.wroc.plot: plot the ROC curve.ci.wauc: confidence intervals for the AUC.ht.indep: hypothesis test for the comparison of two
independent AUCs.ht.paired: hypothesis test for the comparison of two
paired AUCs.The methodology proposed for the above-mentioned functions can be found in the following references:
Iparragirre, A., Barrio, I., Aramendi, J. and Arostegui, I. (2022). Estimation of cut-off points under complex-sampling design data. SORT-Statistics and Operations Research Transactions 46(1), 137–158.
Iparragirre, A., Barrio, I. and Arostegui, I. (2023). Estimation of the ROC curve and the area under it with complex survey data. Stat 12(1), e635.
Iparragirre, A. and Barrio, I. (2024). Optimism Correction of the AUC with Complex Survey Data. In: Einbeck, J., Maeng, H., Ogundimu, E., Perrakis, K. (eds) Developments in Statistical Modelling. IWSM 2024. Contributions to Statistics. Springer, Cham.
To install the package from CRAN:
install.packages("svyROC")To install the most updated version of the package from GitHub run the following code:
devtools::install_github("aiparragirre/svyROC")We need information on three elements for each unit in the sample in
order to estimate the ROC curve (wroc() function) and AUC
(wauc() function):
response.var: variable indicating the dichotomous
response variable.phat.var: predicted probabilities of event.weights.var: variable indicating the sampling
weights.We can put these three vectors in a data frame, or save them
separately in three different vectors. The data set
example_data_wroc is set as an example in the package. We
also need to define the tags for events and non-events.
library(svyROC)
data(example_data_wroc)
mycurve <- wroc(response.var = "y", phat.var = "phat", weights.var = "weights",
data = example_data_wroc,
tag.event = 1, tag.nonevent = 0)
# Or equivalently
mycurve <- wroc(response.var = example_data_wroc$y,
phat.var = example_data_wroc$phat,
weights.var = example_data_wroc$weights,
tag.event = 1, tag.nonevent = 0)Similarly, we can run the following code to estimate the AUC:
auc.obj <- wauc(response.var = "y",
phat.var = "phat",
weights.var = "weights",
tag.event = 1,
tag.nonevent = 0,
data = example_data_wroc)
# Or equivalently
auc.obj <- wauc(response.var = example_data_wroc$y,
phat.var = example_data_wroc$phat,
weights.var = example_data_wroc$weights,
tag.event = 1, tag.nonevent = 0)We can correct the optimism of the weighted estimate of the AUC by
means of replicate weights, as proposed in Iparragirre and Barrio
(2024), by means of the corrected.wauc() function. For this
purpose, we additionally need information on the covariates and the
sampling design. Here is an example of the usage of this function:
data(example_variables_wroc)
mydesign <- survey::svydesign(ids = ~cluster, strata = ~strata,
weights = ~weights, nest = TRUE,
data = example_variables_wroc)
m <- survey::svyglm(y ~ x1 + x2 + x3 + x4 + x5 + x6, design = mydesign,
family = quasibinomial())
phat <- predict(m, newdata = example_variables_wroc, type = "response")
myaucw <- wauc(response.var = example_variables_wroc$y, phat.var = phat,
weights.var = example_variables_wroc$weights)
# Correction of the AUCw:
set.seed(1)
cor <- corrected.wauc(data = example_variables_wroc,
formula = y ~ x1 + x2 + x3 + x4 + x5 + x6,
tag.event = 1, tag.nonevent = 0,
weights.var = "weights", strata.var = "strata", cluster.var = "cluster",
method = "dCV", dCV.method = "pooling", k = 10, R = 20)
# Or equivalently:
set.seed(1)
cor <- corrected.wauc(design = mydesign,
formula = y ~ x1 + x2 + x3 + x4 + x5 + x6,
tag.event = 1, tag.nonevent = 0,
method = "dCV", dCV.method = "pooling", k = 10, R = 20)We can also estimate the sensitivity (wse()) and
specificity (wsp()) parameters for a specific cut-off point
considering sampling weights. For this purpose, we need to indicate the
cut-off point we want to use in the function by means of the argument
cutoff.value:
# Specificity ----------------------------------------------------------
sp.obj <- wsp(response.var = "y",
phat.var = "phat",
weights.var = "weights",
tag.nonevent = 0,
cutoff.value = 0.5,
data = example_data_wroc)
# Or equivalently
sp.obj <- wsp(response.var = example_data_wroc$y,
phat.var = example_data_wroc$phat,
weights.var = example_data_wroc$weights,
tag.nonevent = 0,
cutoff.value = 0.5)
# Sensitivity ----------------------------------------------------------
se.obj <- wse(response.var = "y",
phat.var = "phat",
weights.var = "weights",
tag.event = 1,
cutoff.value = 0.5,
data = example_data_wroc)
# Or equivalently
se.obj <- wse(response.var = example_data_wroc$y,
phat.var = example_data_wroc$phat,
weights.var = example_data_wroc$weights,
tag.event = 1,
cutoff.value = 0.5)Finally, use the function wocp() to obtain optimal
cut-off points for individual classification as proposed in Iparragirre
et al (2022). Some functions of the package
OptimalCutpoints have been modified in order for them to
consider sampling weights:
Lopez-Raton, M., Rodriguez-Alvarez, M.X, Cadarso-Suarez, C. and Gude-Sampedro, F. (2014). OptimalCutpoints: An R Package for Selecting Optimal Cutpoints in Diagnostic Tests. Journal of Statistical Software 61(8), 1–36.
One of the methods proposed in the paper needs to be selected when
running the function by means of the argument method:
Youden, MaxProdSpSe, ROC01 or
MaxEfficiency.
myocp <- wocp(response.var = "y",
phat.var = "phat", weights.var = "weights",
tag.event = 1,
tag.nonevent = 0,
method = "Youden",
data = example_data_wroc)
# Or equivalently
myocp <- wocp(example_data_wroc$y,
example_data_wroc$phat,
example_data_wroc$weights,
tag.event = 1,
tag.nonevent = 0,
method = "Youden")If you want to draw the optimal cut-off point in the ROC curve, then
use the function wroc.plot() and indicate the method by
means of the argument cutoff.method in the function
wroc() as follows:
mycurve <- wroc(response.var = "y",
phat.var = "phat",
weights.var = "weights",
data = example_data_wroc,
tag.event = 1,
tag.nonevent = 0,
cutoff.method = "Youden")
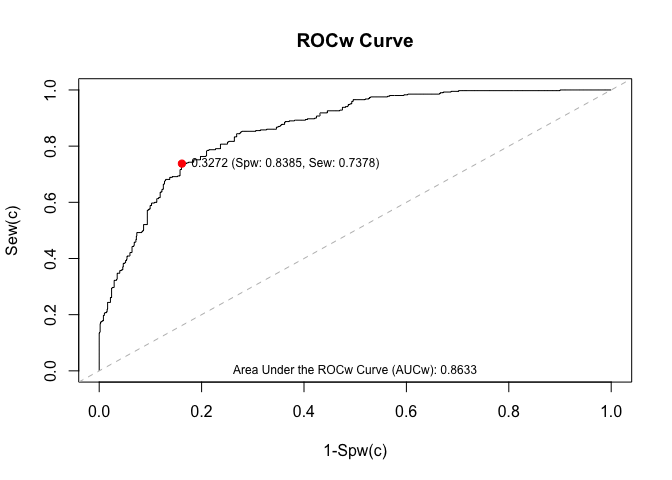
wroc.plot(x = mycurve,
print.auc = TRUE,
print.cutoff = TRUE)
These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.