The hardware and bandwidth for this mirror is donated by METANET, the Webhosting and Full Service-Cloud Provider.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]metanet.ch.
swjm implements stagewise (forward-stepwise) variable
selection for joint models of recurrent events and terminal events
(semi-competing risks). Two model frameworks are supported:
| Model | Type | alpha (first p) |
beta (second p) |
|---|---|---|---|
| JFM | Cox frailty | recurrence/readmission | terminal/death |
| JSCM | Scale-change (AFT) | recurrence/readmission | terminal/death |
Three penalty types are available: cooperative lasso
("coop"), lasso ("lasso"), and group lasso
("group"). The cooperative lasso encourages shared support
between the readmission and death coefficient vectors.
# From the package directory:
devtools::install("swjm")All functions expect a data frame with columns id,
t.start, t.stop, event (1 =
readmission, 0 = terminal/censoring row), status (1 =
death, 0 = alive/censored), and covariate columns
x1, ..., xp.
library(swjm)
# Joint Frailty Model — scenario 1
set.seed(42)
dat_jfm <- generate_data(n = 200, p = 100, scenario = 1, model = "jfm")
Data_jfm <- dat_jfm$data
# JSCM — same scenario
set.seed(42)
dat_jscm <- generate_data(n = 200, p = 100, scenario = 1, model = "jscm")
#> Call:
#> reReg::simGSC(n = n, summary = TRUE, para = para, xmat = X, censoring = C,
#> frailty = gamma, tau = 60)
#>
#> Summary:
#> Sample size: 200
#> Number of recurrent event observed: 343
#> Average number of recurrent event per subject: 1.715
#> Proportion of subjects with a terminal event: 0.175
Data_jscm <- dat_jscm$dataJFM data: n = 200 subjects, 382 readmission events, 48 deaths
fit_jfm <- stagewise_fit(Data_jfm, model = "jfm", penalty = "coop")
fit_jfm
#> Stagewise path (jfm/coop)
#>
#> Covariates (p): 100
#> Iterations: 5000
#> Lambda range: [0.08902, 1.301]
#> Active at final step: 44 readmission, 34 death
#> Readmission (alpha): 1, 2, 3, 4, 5, 7, 9, 10, 12, 14, 15, 17, 23, 33, 35, 37, 38, 39, 41, 43, 47, 49, 52, 55, 57, 60, 62, 64, 66, 67, 69, 71, 73, 74, 77, 79, 80, 82, 92, 93, 95, 96, 97, 98
#> Death (beta): 3, 4, 5, 7, 9, 10, 12, 14, 15, 17, 25, 37, 38, 39, 41, 43, 47, 49, 52, 57, 60, 62, 66, 67, 73, 74, 78, 80, 82, 84, 95, 96, 97, 98The fit object now exposes alpha (readmission) and
beta (death) as separate p × (k+1)
matrices — one column per stagewise step — in addition to the combined
theta matrix (2p × (k+1)):
p <- 100
k_final <- ncol(fit_jfm$alpha)
a_final <- round(fit_jfm$alpha[, k_final], 4)Nonzero alpha entries at final step:
a_final[a_final != 0]
#> [1] 1.0057 -0.9787 0.0183 -0.0577 0.0185 0.0151 0.9614 -0.8484 0.0124
#> [10] 0.0423 -0.0284 0.0199 -0.0020 0.0240 -0.0240 0.0064 0.0781 -0.0061
#> [19] 0.0125 0.0046 -0.0429 0.0396 0.0092 0.0680 -0.0550 -0.0500 -0.0007
#> [28] 0.0200 0.0742 0.0005 -0.0350 0.0370 0.0099 -0.0234 0.0450 0.0790
#> [37] -0.0072 0.0013 -0.0200 0.0150 0.0085 0.0027 -0.0618 0.0272summary() shows a compact table of path-end coefficients
with variable type (shared, readmission-only, or death-only):
summary(fit_jfm)
#> Stagewise path (jfm/coop)
#>
#> p = 100 | 5000 iterations | lambda: [0.08902, 1.301]
#> Decreasing path: 784 steps
#>
#> Path-end coefficients (nonzero variables):
#>
#> Variable alpha beta Type
#> ---------- ---------- ---------- ----------------
#> x9 +0.9614 +0.6629 shared (+)
#> x4 -0.0577 -0.9979 shared (+)
#> x10 -0.8484 -0.7274 shared (+)
#> x2 -0.9787 — readmission only
#> x3 +0.0183 +0.8049 shared (+)
#> x74 -0.0234 -0.3101 shared (+)
#> x1 +1.0057 — readmission only
#> x67 +0.0005 +0.1393 shared (+)
#> x57 -0.0550 +0.1330 shared (–)
#> x38 +0.0781 +0.0857 shared (+)
#> x97 -0.0618 -0.0583 shared (+)
#> x66 +0.0742 +0.0415 shared (+)
#> x41 +0.0125 +0.0937 shared (+)
#> x55 +0.0680 — readmission only
#> x79 +0.0790 — readmission only
#> x60 -0.0500 +0.0070 shared (–)
#> x49 +0.0396 +0.0305 shared (+)
#> x98 +0.0272 +0.0792 shared (+)
#> x15 -0.0284 -0.0639 shared (+)
#> x7 +0.0151 +0.1160 shared (+)
#> x82 +0.0013 +0.0750 shared (+)
#> x5 +0.0185 +0.0527 shared (+)
#> x39 -0.0061 -0.0657 shared (+)
#> x14 +0.0423 +0.0076 shared (+)
#> x47 -0.0429 -0.0135 shared (+)
#> x71 +0.0370 — readmission only
#> x77 +0.0450 — readmission only
#> x69 -0.0350 — readmission only
#> x35 -0.0240 — readmission only
#> x33 +0.0240 — readmission only
#> x17 +0.0199 +0.0020 shared (+)
#> x84 — -0.0790 death only
#> x37 +0.0064 +0.0394 shared (+)
#> x78 — +0.0710 death only
#> x64 +0.0200 — readmission only
#> x92 -0.0200 — readmission only
#> x25 — +0.0200 death only
#> x80 -0.0072 -0.0165 shared (+)
#> x93 +0.0150 — readmission only
#> x12 +0.0124 +0.0085 shared (+)
#> x73 +0.0099 +0.0084 shared (+)
#> x95 +0.0085 +0.0170 shared (+)
#> x52 +0.0092 +0.0092 shared (+)
#> x43 +0.0046 +0.0038 shared (+)
#> x96 +0.0027 +0.0054 shared (+)
#> x23 -0.0020 — readmission only
#> x62 -0.0007 -0.0007 shared (+)
#>
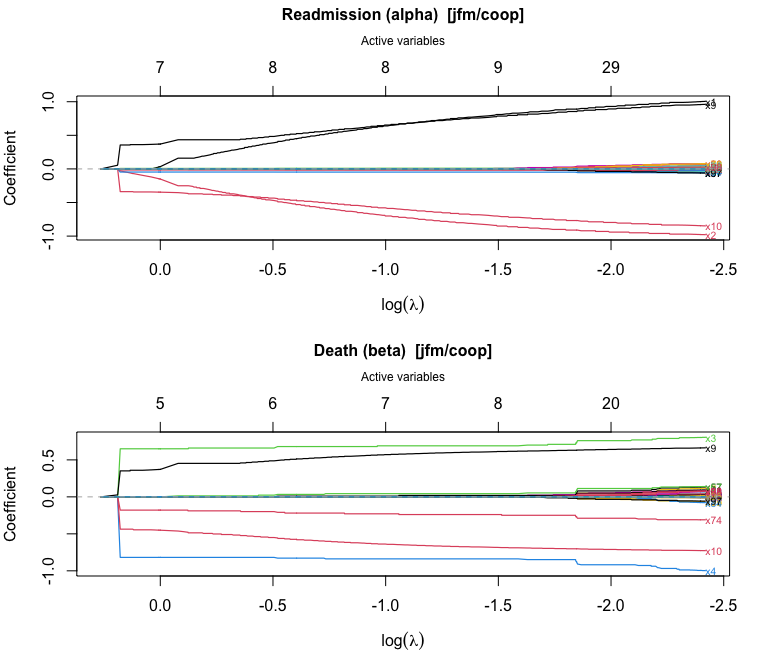
#> Inactive: x6, x8, x11, x13, x16, x18, x19, x20, x21, x22, x24, x26, x27, x28, x29, x30, x31, x32, x34, x36, x40, x42, x44, x45, x46, x48, x50, x51, x53, x54, x56, x58, x59, x61, x63, x65, x68, x70, x72, x75, x76, x81, x83, x85, x86, x87, x88, x89, x90, x91, x94, x99, x100plot() produces a glmnet-style coefficient trajectory
plot with the number of active variables on the top axis:
plot(fit_jfm)
cv_stagewise() evaluates a cross-fitted
estimating-equation score norm over a grid of lambda values.
lambda_path <- fit_jfm$lambda
dec_idx <- swjm:::extract_decreasing_indices(lambda_path)
lambda_seq <- lambda_path[dec_idx]
cv_jfm <- cv_stagewise(
Data_jfm, model = "jfm", penalty = "coop",
lambda_seq = lambda_seq, K = 3L
)
cv_jfm
#> Cross-validation (jfm/coop)
#>
#> Covariates (p): 100
#> Lambda grid size: 784
#> Best position (combined): 438 (lambda = 0.9241)
#> Selected variables: 8 readmission, 6 death
#> Readmission (alpha): 1, 2, 3, 4, 9, 10, 67, 74
#> Death (beta): 3, 4, 9, 10, 67, 74The CV object stores alpha and beta at the
optimal lambda, plus n_active_alpha,
n_active_beta, and n_active variable-count
vectors across the full lambda grid.
plot(cv_jfm)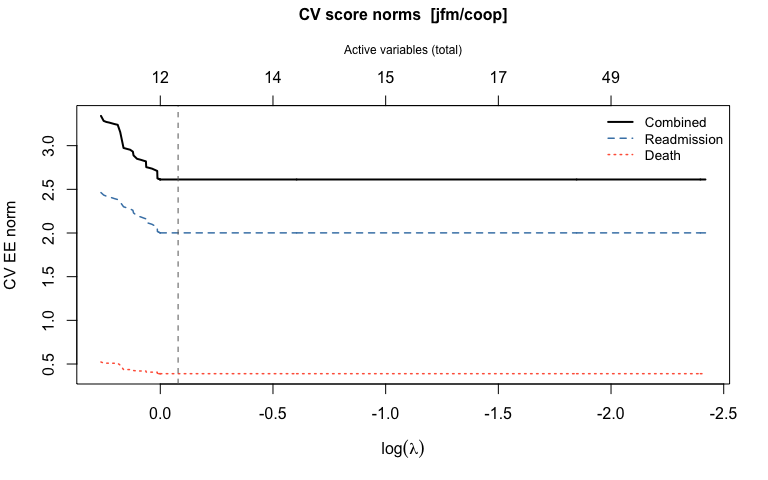
# summary() shows a formatted table of selected coefficients
summary(cv_jfm)
#> CV-selected model (jfm/coop)
#>
#> p = 100 | Lambda grid: 784 steps | CV optimal: step 438 (lambda = 0.9241)
#>
#> Selected coefficients (8 readmission, 6 death):
#>
#> Variable alpha beta Type
#> ---------- ---------- ---------- ----------------
#> x9 +0.4338 +0.4528 shared (+)
#> x4 -0.0466 -0.8184 shared (+)
#> x10 -0.3531 -0.4622 shared (+)
#> x3 +0.0075 +0.6503 shared (+)
#> x2 -0.2507 — readmission only
#> x74 -0.0162 -0.1793 shared (+)
#> x1 +0.1597 — readmission only
#> x67 +0.0004 +0.0133 shared (+)
#>
#> Inactive (92): x5, x6, x7, x8, x11, x12, x13, x14, x15, x16, x17, x18, x19, x20, x21, x22, x23, x24, x25, x26, x27, x28, x29, x30, x31, x32, x33, x34, x35, x36, x37, x38, x39, x40, x41, x42, x43, x44, x45, x46, x47, x48, x49, x50, x51, x52, x53, x54, x55, x56, x57, x58, x59, x60, x61, x62, x63, x64, x65, x66, x68, x69, x70, x71, x72, x73, x75, x76, x77, x78, x79, x80, x81, x82, x83, x84, x85, x86, x87, x88, x89, x90, x91, x92, x93, x94, x95, x96, x97, x98, x99, x100Direct access to selected coefficients:
# coef() returns the combined numeric vector c(alpha, beta) for compatibility
theta_best <- coef(cv_jfm)baseline_hazard() evaluates the cumulative baseline
hazards at any desired time points (Breslow for JFM; Nelson-Aalen on the
accelerated scale for JSCM):
bh <- baseline_hazard(cv_jfm, times = c(0.5, 1.0, 2.0, 4.0, 6.0))
print(bh)
#> time cumhaz_readmission cumhaz_death
#> 1 0.5 0.8302786 0.04821411
#> 2 1.0 1.3776457 0.06859515
#> 3 2.0 1.9078200 0.12461276
#> 4 4.0 3.1933429 0.28725353
#> 5 6.0 4.5481622 0.33588232predict() computes subject-specific readmission-free and
death-free survival curves, together with the per-predictor
contributions to each linear predictor:
# Three hypothetical new subjects (covariate vectors of length p = 100)
set.seed(7)
newz <- matrix(rnorm(300), nrow = 3, ncol = 100)
colnames(newz) <- paste0("x", 1:100)
pred <- predict(cv_jfm, newdata = newz)
pred
#> swjm predictions (jfm)
#>
#> Subjects: 3
#> Time points: 430
#> Time range: [0.0002844, 6.281]
#>
#> Use plot() to visualize survival curves and predictor contributions.
# S_re: readmission-free survival — first 10 time points
round(pred$S_re[, 1:10], 3)
#> t=0.0002844 t=0.0009392 t=0.001171 t=0.001381 t=0.001936 t=0.002432
#> Subject1 0.992 0.983 0.983 0.974 0.966 0.957
#> Subject2 0.994 0.987 0.987 0.980 0.973 0.967
#> Subject3 0.994 0.987 0.987 0.981 0.974 0.967
#> t=0.004535 t=0.004809 t=0.006178 t=0.006205
#> Subject1 0.949 0.940 0.932 0.923
#> Subject2 0.960 0.953 0.947 0.940
#> Subject3 0.961 0.954 0.948 0.941
# Predictor contributions for subject 1 — show only nonzero entries
contrib1 <- round(pred$contrib_re[1, ], 3)
contrib1[contrib1 != 0]
#> x1 x2 x3 x4 x9 x10 x74
#> 0.365 0.103 0.006 -0.102 0.552 -0.209 0.007plot() on a swjm_pred object draws four
panels: survival curves for both processes (all subjects in grey,
highlighted subject in color) plus bar charts of predictor contributions
for the selected subject.
plot(pred, which_subject = 1)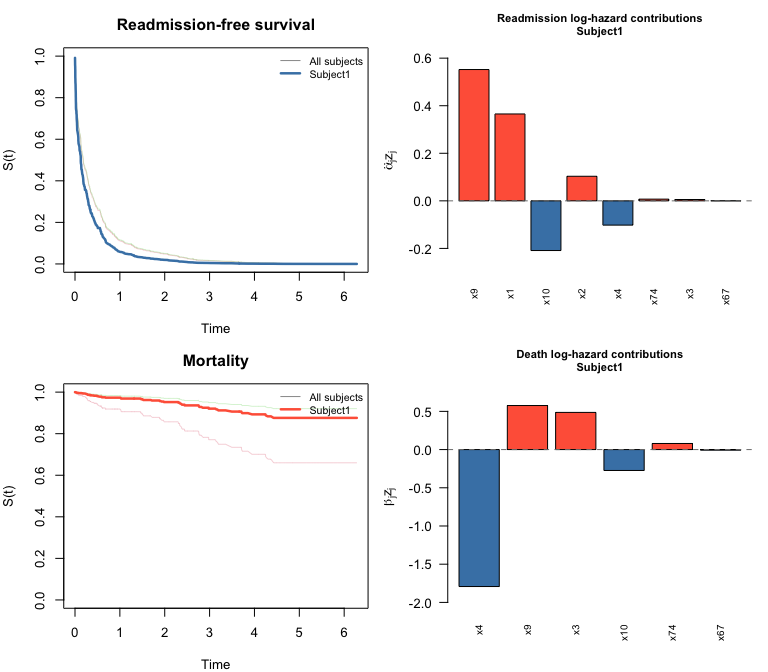
The same workflow applies to the JSCM. baseline_hazard()
and predict() now work for both models: for JSCM, survival
curves are estimated via a Nelson-Aalen baseline on the accelerated time
scale. In the AFT interpretation, \(e^{\hat\alpha^\top z_i}\) is the
time-acceleration factor for subject \(i\): greater than 1 means events happen
sooner, less than 1 means later.
fit_jscm <- stagewise_fit(Data_jscm, model = "jscm", penalty = "coop")
lambda_path_jscm <- fit_jscm$lambda
dec_idx_jscm <- swjm:::extract_decreasing_indices(lambda_path_jscm)
lambda_seq_jscm <- lambda_path_jscm[dec_idx_jscm]
cv_jscm <- cv_stagewise(
Data_jscm, model = "jscm", penalty = "coop",
lambda_seq = lambda_seq_jscm, K = 3L
)
summary(cv_jscm)
#> CV-selected model (jscm/coop)
#>
#> p = 100 | Lambda grid: 30 steps | CV optimal: step 3 (lambda = 0.6838)
#>
#> Selected coefficients (1 readmission, 1 death):
#>
#> Variable alpha beta Type
#> ---------- ---------- ---------- ----------------
#> x10 -0.0180 -0.0088 shared (+)
#>
#> Inactive (99): x1, x2, x3, x4, x5, x6, x7, x8, x9, x11, x12, x13, x14, x15, x16, x17, x18, x19, x20, x21, x22, x23, x24, x25, x26, x27, x28, x29, x30, x31, x32, x33, x34, x35, x36, x37, x38, x39, x40, x41, x42, x43, x44, x45, x46, x47, x48, x49, x50, x51, x52, x53, x54, x55, x56, x57, x58, x59, x60, x61, x62, x63, x64, x65, x66, x67, x68, x69, x70, x71, x72, x73, x74, x75, x76, x77, x78, x79, x80, x81, x82, x83, x84, x85, x86, x87, x88, x89, x90, x91, x92, x93, x94, x95, x96, x97, x98, x99, x100set.seed(7)
newz_jscm <- matrix(runif(600, -1, 1), nrow = 3, ncol = 100)
#> Warning in matrix(runif(600, -1, 1), nrow = 3, ncol = 100): data length differs
#> from size of matrix: [600 != 3 x 100]
pred_jscm <- predict(cv_jscm, newdata = newz_jscm)Recurrence time-acceleration factors:
round(pred_jscm$time_accel_re, 3)
#> Subject1 Subject2 Subject3
#> 1.000 0.983 1.005plot() draws the same four-panel layout as for JFM:
survival curves for both processes plus bar charts of log
time-acceleration contributions.
plot(pred_jscm, which_subject = 1)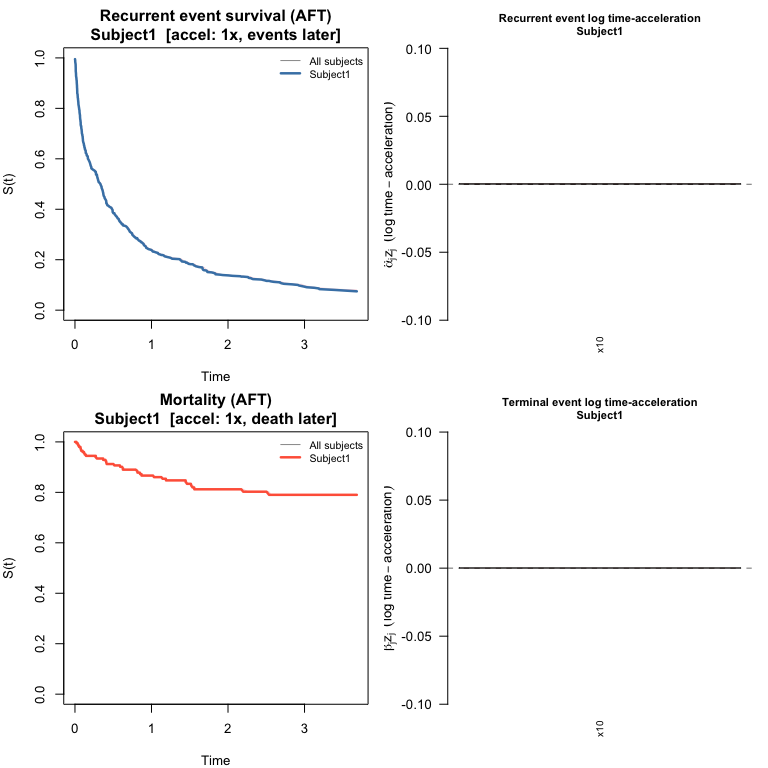
fit_lasso <- stagewise_fit(Data_jfm, model = "jfm", penalty = "lasso")
cv_lasso <- cv_stagewise(Data_jfm, model = "jfm", penalty = "lasso", K = 3L)
summary(cv_lasso)
fit_group <- stagewise_fit(Data_jfm, model = "jfm", penalty = "group")
cv_group <- cv_stagewise(Data_jfm, model = "jfm", penalty = "group", K = 3L)
summary(cv_group)Compare CV-optimal estimates to the true generating coefficients. Variables that are truly nonzero or were selected are shown; all others were correctly excluded.
p <- 100
# JFM: variables of interest (true signal or selected)
show_jfm <- sort(which(dat_jfm$alpha_true != 0 | cv_jfm$alpha != 0 |
dat_jfm$beta_true != 0 | cv_jfm$beta != 0))
coef_df <- data.frame(
variable = paste0("x", show_jfm),
true_alpha = round(dat_jfm$alpha_true[show_jfm], 3),
est_alpha = round(cv_jfm$alpha[show_jfm], 3),
true_beta = round(dat_jfm$beta_true[show_jfm], 3),
est_beta = round(cv_jfm$beta[show_jfm], 3)
)
colnames(coef_df) <- c("variable", "alpha_true", "alpha_est", "beta_true", "beta_est")
print(coef_df, row.names = FALSE)
#> variable alpha_true alpha_est beta_true beta_est
#> x1 1.1 0.160 0.1 0.000
#> x2 -1.1 -0.251 -0.1 0.000
#> x3 0.1 0.007 1.1 0.650
#> x4 -0.1 -0.047 -1.1 -0.818
#> x9 1.0 0.434 1.0 0.453
#> x10 -1.0 -0.353 -1.0 -0.462
#> x67 0.0 0.000 0.0 0.013
#> x74 0.0 -0.016 0.0 -0.179JFM alpha: TP=6 FP=2 FN=0 | beta: TP=4 FP=2 FN=2
show_jscm <- sort(which(dat_jscm$alpha_true != 0 | cv_jscm$alpha != 0 |
dat_jscm$beta_true != 0 | cv_jscm$beta != 0))
coef_jscm <- data.frame(
variable = paste0("x", show_jscm),
true_alpha = round(dat_jscm$alpha_true[show_jscm], 3),
est_alpha = round(cv_jscm$alpha[show_jscm], 3),
true_beta = round(dat_jscm$beta_true[show_jscm], 3),
est_beta = round(cv_jscm$beta[show_jscm], 3)
)
colnames(coef_jscm) <- c("variable", "alpha_true", "alpha_est", "beta_true", "beta_est")
print(coef_jscm, row.names = FALSE)
#> variable alpha_true alpha_est beta_true beta_est
#> x1 1.1 0.000 0.1 0.000
#> x2 -1.1 0.000 -0.1 0.000
#> x3 0.1 0.000 1.1 0.000
#> x4 -0.1 0.000 -1.1 0.000
#> x9 1.0 0.000 1.0 0.000
#> x10 -1.0 -0.018 -1.0 -0.009JSCM alpha: TP=1 FP=0 FN=5 | beta: TP=1 FP=0 FN=5
We use the timeROC package (Blanche et al., 2013) to
compute cause-specific time-varying AUC in the competing-risk framework.
Each subject contributes at most a first-readmission event (cause 1) and
a death event (cause 2). Each sub-model is assessed with its own linear
predictor: \(\hat\alpha^\top z_i\) for
readmission, \(\hat\beta^\top z_i\) for
death.
Note: AUC is evaluated on the training data for illustration. In practice use held-out or cross-validated predictions.
# Construct competing-risk dataset:
# Keep first readmission (event==1 & t.start==0) + death/censor (event==0).
# Status: 1 = first readmission, 2 = death, 0 = censored.
.cr_data <- function(Data) {
d3 <- Data[Data$event == 0 | (Data$event == 1 & Data$t.start == 0), ]
d3 <- d3[order(d3$id, d3$t.start, d3$t.stop), ]
status <- ifelse(d3$event == 1 & d3$status == 0, 1L,
ifelse(d3$event == 0 & d3$status == 0, 0L, 2L))
list(data = d3, status = status)
}
cr_jfm <- .cr_data(Data_jfm)
cr_jscm <- .cr_data(Data_jscm)
# Baseline covariates (one row per subject)
Z_jfm <- as.matrix(Data_jfm[!duplicated(Data_jfm$id), paste0("x", 1:p)])
Z_jscm <- as.matrix(Data_jscm[!duplicated(Data_jscm$id), paste0("x", 1:p)])
# Markers expanded to row level: alpha^T z for readmission, beta^T z for death
M_re_jfm <- drop(Z_jfm %*% cv_jfm$alpha)[cr_jfm$data$id]
M_de_jfm <- drop(Z_jfm %*% cv_jfm$beta)[cr_jfm$data$id]
M_re_jscm <- drop(Z_jscm %*% cv_jscm$alpha)[cr_jscm$data$id]
M_de_jscm <- drop(Z_jscm %*% cv_jscm$beta)[cr_jscm$data$id]if (!requireNamespace("timeROC", quietly = TRUE))
install.packages("timeROC")
library(survival)
library(timeROC)
# Evaluation grid: 20 points spanning the 10th-85th percentile of event times
.tgrid <- function(t_vec, status, n = 20) {
t_ev <- t_vec[status > 0]
seq(quantile(t_ev, 0.10), quantile(t_ev, 0.85), length.out = n)
}
t_jfm <- .tgrid(cr_jfm$data$t.stop, cr_jfm$status)
t_jscm <- .tgrid(cr_jscm$data$t.stop, cr_jscm$status)
# Readmission AUC: alpha^T z marker, cause = 1
roc_re_jfm <- timeROC(T = cr_jfm$data$t.stop, delta = cr_jfm$status,
marker = M_re_jfm, cause = 1, weighting = "marginal",
times = t_jfm, ROC = FALSE, iid = FALSE)
roc_re_jscm <- timeROC(T = cr_jscm$data$t.stop, delta = cr_jscm$status,
marker = M_re_jscm, cause = 1, weighting = "marginal",
times = t_jscm, ROC = FALSE, iid = FALSE)
# Death AUC: beta^T z marker, cause = 2
roc_de_jfm <- timeROC(T = cr_jfm$data$t.stop, delta = cr_jfm$status,
marker = M_de_jfm, cause = 2, weighting = "marginal",
times = t_jfm, ROC = FALSE, iid = FALSE)
roc_de_jscm <- timeROC(T = cr_jscm$data$t.stop, delta = cr_jscm$status,
marker = M_de_jscm, cause = 2, weighting = "marginal",
times = t_jscm, ROC = FALSE, iid = FALSE).get_auc <- function(roc, cause) {
auc <- roc[[paste0("AUC_", cause)]]
if (is.null(auc)) auc <- roc$AUC
if (is.null(auc) || !is.numeric(auc)) return(rep(NA_real_, length(roc$times)))
if (length(auc) == length(roc$times) + 1) auc <- auc[-1]
as.numeric(auc)
}
old_par <- par(mfrow = c(1, 2), mar = c(4.5, 4, 3, 1))
plot(t_jfm, .get_auc(roc_re_jfm, 1), type = "l", lwd = 2, col = "steelblue",
xlab = "Time", ylab = "AUC(t)", main = "JFM", ylim = c(0.4, 1))
lines(t_jfm, .get_auc(roc_de_jfm, 2), lwd = 2, col = "tomato", lty = 2)
abline(h = 0.5, lty = 3, col = "grey60")
legend("bottomleft", c("Readmission", "Death"),
col = c("steelblue", "tomato"), lwd = 2, lty = c(1, 2),
bty = "n", cex = 0.85)
plot(t_jscm, .get_auc(roc_re_jscm, 1), type = "l", lwd = 2, col = "steelblue",
xlab = "Time", ylab = "AUC(t)", main = "JSCM", ylim = c(0.4, 1))
lines(t_jscm, .get_auc(roc_de_jscm, 2), lwd = 2, col = "tomato", lty = 2)
abline(h = 0.5, lty = 3, col = "grey60")
legend("bottomleft", c("Readmission", "Death"),
col = c("steelblue", "tomato"), lwd = 2, lty = c(1, 2),
bty = "n", cex = 0.85)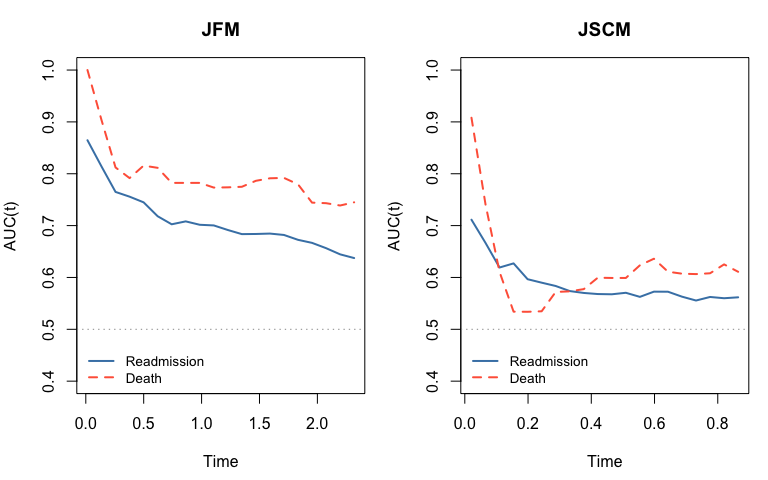
par(old_par)theta is always stored as
c(alpha, beta): theta[1:p] =
alpha (readmission), theta[(p+1):(2p)] =
beta (death). This holds for both the JFM and JSCM.stagewise_fit() also returns alpha and
beta as separate p × (k+1) matrices for
convenient access to the full regularization path.cv_stagewise() stores alpha and
beta at the CV-optimal lambda, plus variable-count vectors
n_active_alpha, n_active_beta,
n_active.cv$baseline and accessible via
baseline_hazard(cv, times). Works for both models: Breslow
estimator for JFM; Nelson-Aalen on the accelerated time scale for JSCM.
Used internally by predict().predict(cv, newdata) works for both models and returns a
swjm_pred object with S_re, S_de
(survival matrices), linear predictors lp_re,
lp_de, and predictor-contribution matrices
contrib_re, contrib_de (\(\hat\alpha_j z_{ij}\) and \(\hat\beta_j z_{ij}\)).
time_accel_re and
time_accel_de (\(e^{\hat\alpha^\top z_i}\), \(e^{\hat\beta^\top z_i}\)) — the
multiplicative factors by which each subject’s event times are scaled
relative to baseline. Contributions \(\hat\alpha_j z_{ij}\) are log
time-acceleration contributions: \(e^{\hat\alpha_j z_{ij}} > 1\) shortens
event times; \(< 1\) lengthens
them.(alpha_j, beta_j) with the L2 norm when signs agree,
and L1 when they disagree — encouraging variables that affect both
processes to enter together with the same sign.These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.